Sensitive miRNA markers for the detection and management of NSTEMI acute myocardial infarction patients
Introduction
Cardiovascular disease (CVD) is a chronic ailment that affects millions worldwide (1). It is a significant cause of mortality, with a worsening trend in developing countries (2). In China, the mortality rates approximate 280 per 100,000 people in 2014 (3). It is the leading cause of death in China in 2014 with cancer deaths a distant second (3). A correct diagnosis and prognosis is important that may provide better immediate care. For instance, acute myocardial infarction (AMI) patients can be placed on immediate initiation of reperfusion therapy to restore blood flow. Creatine kinase-MB (CK-MB) isoenzymes and cardiac troponins are commonly used biomarkers for detection and treatment monitoring (4). These markers have demonstrated clear clinical utility to help manage the disease (5). However, in certain cases such as geriatric patients with non-ST-elevation MI (NSTEMI), these tests are often challenging to interpret (6). The exploration of new biomarkers that can provide sensitive and real time probing to the acute condition is necessary.
Several classes of circulating biomarkers within plasma demonstrated strong potential to fill in the gaps in disease detection and prognosis (7). These include circulating microRNAs (miRNAs) (8), mitochondrial DNA (9) and nuclear DNA (10) that show strong significance to different stages of CVD. The main advantage is faster response time upon disease onset compared with existing markers such as conventional cardiac troponins. Both cardiac troponin T (cTnT) and cardiac troponin I (cTnI) show significant elevated levels only around 6 hours, and even with newer generation of high sensitive immunoassays, it is still difficult to diagnose early AMI (7). In comparison, nucleotide based biomarkers in blood have been observed to be detectable after 1 hour (11), potentially allowing quicker diagnosis of the condition. In prior studies, several miRNAs have been proposed such as miR-1 (12) and miR-133 (13), which are cardiac-specific markers. Other studies point to the usefulness of miR-208 and miR-499 in the disease management of CVD (14). This forms the basis of our current study and the selection of suitable miRNAs in our patient cohort.
In the current study, we hypothesize that miRNAs can play an essential role in early detection and patient monitoring. We are further interested to address if variations in miRNA levels correlate to secondary complications, given the short half-life of these molecules. miRNAs are small size non-coding RNA and its biological functions are not fully understood (15). Previous studies have implicated its presence to cellular regulatory functions such as cell proliferation, differentiation and program cell death (16,17). Certain miRNAs are cardiac specific (18) and preliminary studies highlight their critical roles in development and function (19,20). The main objective of the current study is to examine the miRNA levels in NSTEMI patients during disease onset and therapy. Serial sampling is conducted to understand the markers’ variations during disease development.
Methods
Patient groups and disease characteristics
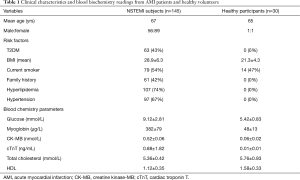
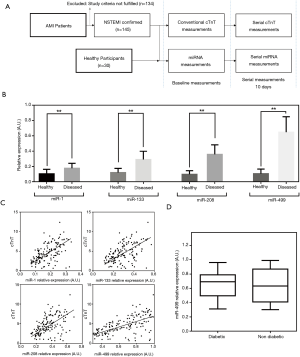
For this study, a total of 145 patients were recruited. These patients had presentation of AMI and were NSTEMI cases. The study design process is outline in Figure 1A. Patients were admitted to the First Affiliated Hospital of Lanzhou University and provided written informed consent to be part of the study. All procedures regarding patient recruitment and sample processing were approved by the institutional review board (IRB, 2543/00135). All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards. The details and risk factors associated with these participants are presented in Table 1. The mean age of the cohort was 67 (95% CI, 63–71) years old. NSTEMI was diagnosed following guidelines of the American College of Cardiology (21). Briefly, the study patients were presented with acute ischaemic-type chest pain and clinically diagnosed by biochemical markers. Adjudication of the final diagnosis was provided by the attending cardiologist. Patients with renal disease were excluded from this study due to the influence on cardiac troponin levels. To ensure a better comparison, 30 healthy volunteers with a similar mean age of 65 years old were recruited. All healthy volunteers had normal electrocardiographic findings and with no history of CVD. The gender ratio was 1:1 (male:female). Most healthy volunteers were recruited during annual health check-ups at the hospital.

Full table
Sample collection schedule and blood processing
The first blood sampling was conducted 2–4 hours upon admission and both miRNA levels and troponin T measurements were conducted. This ensured a fair comparison among two distinct parameters. For serial measurements, a total of 5 mL of blood was taken each day for a total of 10 days for collection of miRNAs and cardiac troponin levels. To recover plasma, samples were spun at 10,000 g in a high-speed centrifuge for 10 minutes at 4 °C. The supernatant was carefully removed to a new centrifuge tube and spun down again using the same setting to remove any contaminating cellular debris. Plasma was split into 2 tubes for miRNA detection and the other for biochemical analysis. Plasma for miRNA detection was processed immediately to convert to cDNA before storage. Samples were stored at −20 °C prior to further processing.
Quantification of miRNAs in plasma and biochemical processing for cardiac troponin
The current study examined a number of miRNAs from clinical blood and healthy volunteers. These included miR-1, miR-133, miR-208 and miR-499. As a first step, small RNA fragments were recovered from plasma specimens at different time points using the mirVana miRNA Isolation Kit (Ambion, Thermo Fisher Scientific, USA). The processing steps strictly following manufacturer’s instructions to obtain small RNAs lesser than 200 nucleotides. For reverse transcription, the miScript II RT Kit (Qiagen Inc. USA) was used for cDNA generation. RT-PCR was performed on the Applied Biosystems VeritiTM Thermal Cycler (Thermo Fisher Scientific, USA). Samples were stored before batch processing for measuring the different miRNA levels. For detection of miRNA, quantitative PCR (qPCR) was performed using primers procured from the miScript Primer Assay (Qiagen Inc., USA) and qPCR reagents using the miScript SYBR Green PCR Kit (Qiagen Inc., USA). qPCR was performed on the Applied Biosystems 7500 system (Thermo Fisher Scientific, USA). For cardiac troponin T (cTnT) measurements, this was performed on the fourth generation assay using the Elecsys® Troponin T high sensitive (Roche Diagnostics, USA).
Statistical analysis
All data were single blinded and laboratory personnel performing the assays were not provided with the clinical details of the patients. The results were analysed using MedCalc (IBM, USA) or Prism 6.0 (GraphPad, USA). Differences among the study groups at baseline were compared using a Student’s t-test to gauge clinical relevance of miRNA measurements. Receiver operating characteristics (ROC) curves were generated to assess the accuracy of each miRNA. Correlation with cardiac troponins was calculated using the Spearman’s rho correlation coefficient. A one-way ANOVA with repeated measures was computed for serial comparisons of miRNA levels in AMI patients. All variables are represented as mean and standard deviation unless otherwise specified. P values of less than 0.05 in each statistical test were considered significant.
Results
Study design, patient characteristics and baseline miRNAs levels
A total of 145 AMI patients were enrolled for the study. The gender ratio was 56:89 (male:female). Another 30 healthy volunteers were recruited to provide a direct comparison of miRNA levels with AMI patients. Details of sample collections and comparisons are outlined in Figure 1A. For all patients, NSTEMI was confirmed and was the target focus group for this study. The study design captured the baseline miRNA levels shortly upon admission and consecutively for 10 days to chart out the variations during disease treatment. Table 1 shows the patients’ characteristics and baseline clinical data upon admission. Among the patient group, 43% were diabetics, average BMI was 28.9 and 42% had family history of CVD. For healthy volunteers, a comprehensive physical examination coupled with a health questionnaire ensured that patients were relatively at low risk of CVD and were non-diabetics.
The baseline miRNA levels are shown in Figure 1B. NSTEMI patient results were presented with significantly higher levels than healthy volunteers for all four different miRNAs using an unpaired Student’s t-test. It was observed that the fold changes for AMI patients varies across different miRNA measurements. Among the four measurements, miR-1 registered the least amount of change and mean average differences were 1.69 folds higher (P<0.001). The largest change was observed in miR-499 quantities with mean average difference of over 6.03 folds (P<0.001). For miR-133 and miR-208, the differences between AMI patients and healthy controls were 2.40 folds (P<0.0001) and 3.63 (P<0.0001) respectively. These results indicated the strong clinical relevance of miRNA measurements to the disease. Comparing with conventional cardiac troponin measurements at baseline, we observed significant correlation with three of four different measurements (Figure 1C). Spearman’s correlation coefficients for miR-1, miR-133, miR-208 and miR-499 were 0.572, 0.511, 0.496 and 0.605 respectively. Among the four parameters, miR-499 was most significantly associated with cTnT measurements for the current study population. To better ascertain the clinical relevance of miRNAs, potential confounding factors were checked against. Based on the patient background, it was observed that a significant portion of the cohort were diabetics (43%). The analysis of miRNA levels was split between patients with diabetes mellitus and those without. Figure 1D showed that there was insignificant statistical evidence to conclude that diabetes had a critical role in these AMI patients. Using a Student’s t-test, there was insignificant evidence to conclude any difference in relative mean detected miR-499 quantities in these two groups (P=0.333).
ROC curve analysis of miRNAs
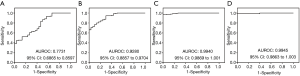
To evaluate the diagnostic accuracy of miRNA used in our study of NSTEMI patients, ROC curves were plotted and compared with the corresponding cTnT derived curves. Figure 2 shows the results of the analysis for different miRNA markers. Of the four different analysis, we observed that miR-499 was most significant. For comparison of studies using the cTnT-hs test, we observed three of four assays demonstrated higher AUROC values comparing with cTnT. Figure 2A shows the direct comparison of miR-1 with a similar AUROC of 0.7731 compared to 0.778 for cTnT-hs (P<0.05). Using miR-133, the associated AUROC (Figure 2B, 0.9280, P>0.05) was much higher than the cTnT derived value. miR-208 and miR-499 had similar AUROC in our cohort analyses. The AUROC calculated from the miR-208 assay was 0.9940 (P<0.05) as shown in Figure 2C. For miR-499 result (Figure 2D), the AUROC measurement was 0.9945 (P<0.05). These results are suggestive of the ability of miRNAs in the diagnosis of AMI for NSTEMI patients with most assays showing diagnostic accuracy superior than the conventional cTnT immunoassay.

Serial measurements of miRNAs for AMI patients
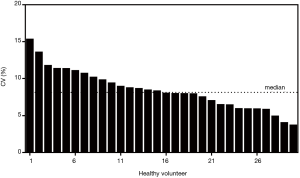
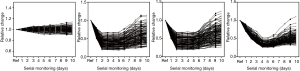
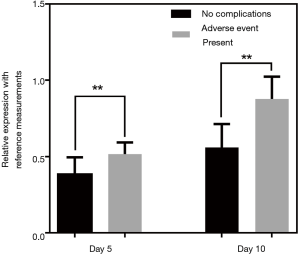
The quantities of each individual miRNA were collated for a consecutive 10 days to gauge the trends during disease management. As a negative control, the same analysis was conducted on the healthy volunteers to gauge the stability of the miRNA levels. Figure 3 shows the plot of the coefficient of variations (CV) for the healthy participants for the 10-day measurements. Median CV value for healthy individuals was 8.14%. Maximum value was 15.3%. These values were fairly low and indicated stable readings in the healthy group for this short-term measurement. For miRNA trending characteristics in NSTEMI patients, there were distinct differences. miR-133 serial analysis showed a relatively uniform profile over the course of 10 days among all 145 patients (Figure 4A). Repeated measures ANOVA indicated insignificant changes in the values for the patient group (P=0.684). For miR-1, miR-208 and miR-499, initial dip in levels were observed after treatment were administered. The lowest mean levels for miR-1 (Figure 4B) and miR-499 (Figure 4C) were seen on day 2. The lowest mean level for miR-208 was registered on day 4 (Figure 4D). Interestingly, a number of patients experienced a spike in miR-499 levels post day 5 and mean levels were monotonic increasing thereafter. Patients were subsequently monitored for any post-AMI complications as part of routine clinical management and it was observed that a significant number who experienced one or more adverse events. These included ischemic complications (angina and reinfarction) and arrhythmic complications. Reinfarction was defined as angina or anginal equivalent accompanied by a rise in serial high sensitivity troponin T measurements. Interestingly, different trends in miRNA levels were seen in patients with different clinical outcomes. The patient group was split into two subgroups based on whether patients had any post-AMI complications and relative levels of miR-499 analyzed on different days. Figure 5 shows the comparison at day 5 and day 10. The results indicated that on both days, the patient subgroup without any post-disease complications were markedly lower for levels in miR-499. Statistical results were performed using Student’s t-tests and indicated a lower P value at day 10. Potentially, this could be useful to identify high risk individuals susceptible to further complications post-AMI.

Discussion
It is of interest to address other forms of potential biomarkers suitable for diagnosis and prognosis of cardiac events. In particular, the study’s intent is to address the levels of miRNAs in NSTEMI patients. The symptoms and conventional disease measurements associated with NSTEMI may be atypical, especially in geriatric patients (6). This can be due to external devices (cardiac pacemaker) or diffused ECG changes as a result of chronic ischemic cardiomyopathy (22,23). A total of 145 subjects were involved to assess the clinical value of four different miRNAs and were monitored for a 10-day period to track the trends associated with the disease. NSTEMI cases are of particular interest due to the insensitivity of conventional cardiac troponin tests (24). In our patient cases, after diagnosis was made of NSTEMI, majority received antiplatelet therapy before further assessment of individual risks. Of the 145 patients, 58 were considered high risk and coronary angiography was carried out and percutaneous coronary intervention (PCI) was performed if required. miRNAs are also advantages as a disease parameter as their expected half-life in the body’s circulatory system is short (25) and may be able to show instantaneous changes during disease development. Our results demonstrated good correlations to cTnT high sensitive test, which is currently one of the main routine assays used to detect MI events. Based on various clinical reports, cTnT high sensitive tests outperforms conventional cTnT (4th generation) assays (26-28) and our study established that miRNA were comparable if not superior in diagnostic accuracy. Serial measurements were also of particular attention as increased levels of miRNA levels appeared to be correlated to secondary complications after AMI onset. This assay potentially can complement current disease management routines to better identify patients with undesirable outcomes.
At baseline reference measurements for NSTEMI patients, we observed elevated levels in all four miRNAs compared to healthy volunteers. This was critical as it established a clear clinical association of these markers to the disease for this particular patient group. Our results were consistent with several other studies in measuring various patient groups with different miRNAs (12,13,29,30). Ai et al. addressed the diagnostic performance of miR-1 in 159 admitted patients with a AUROC curve of 0.85 (12). Kuwabara et al. revealed a strong association of myocardial damage with levels of miR-133 (13). Olivieri et al. discovered that miR-499 can be an effective biomarker for geriatric patients (30). In addition, the miRNAs selected in our panel are mostly cardiac specific. In prior studies, miR-1 and miR-133 had been demonstrated to play critical roles in development and cardiac functions (20,31,32). This makes miRNAs an attractive specific target in the management of CVD. In our study, we observed at least 1.69-fold changes with miR-1 and the maximum fold change was associated to miR-499. Furthermore, we demonstrated that this was not due to other significant diseases like diabetes mellitus. Potentially the diagnostic performance of miRNA in NSTEMI patients could be specific and our ROC analysis showed miR-499 had the highest value of AUROC among the four different candidates. This also closely matched cTnT-hs assays values with a significant correlation coefficient. In our temporal measurements, we observed significant fluctuations in readings for 3 of 4 miRNAs tested whereas values for cardiac troponins remain relatively constant throughout. We postulate that the changes that were captured could be a result of disease development. Thus, besides providing critical information on disease severity and diagnosis, further monitoring of the parameter could be useful in the management of the disease.
Clearance of miRNAs from the circulatory system is fairly rapid from previous studies (25) and hence may be a good indicator for the real time changes in the disease. Of the four tested parameters, miR-133 serial sampling trend matches that of cTnT and showed insignificant variability within these 10 days. This can potentially be useful in cases where cardiac troponin measurements may be hindered such as in geriatric patients. Among the other three miRNA parameters, we observed fairly interesting trends. In the same sample measurement, we noticed that the rate of decrease in the other three miRNA levels varies and miR-133 was the least responsive. This could be related to the state of the disease and would warrant a larger mechanism study in the future to address the molecular functions of these miRNAs. Our study demonstrated clear clinical benefits in short term monitoring for these patients. Given that miR-499 had the largest difference in measured levels during disease onset and after treatment, we used this parameter to correlate to secondary complications that occurred in a subset of patients. It was statistically significant at day 5 that the patient subgroup without complications had lower mean miR-499 levels. At day 10, we observed that difference in levels were higher and the P value comparing both subgroups of patients were lower. This potentially can be a useful tool to address the difficulties in identifying patients at high risk of complications arising from the MI event. Interesting, similar deductions on other patient groups suffering from AMI was noted in plasma circulating DNA levels where changes in absolute levels were a good indicator to identify high risk individuals. It will be interesting to further link these molecular changes to the development of complications that are associated with MI. One limitation of the current study is the lack of comparison with possibly chest pain patients without AMI to address its sensitivity and specificity in detecting infarction for patients in the emergency department. We had compared with healthy volunteers in this study to establish baseline levels in individuals without clear indications of CVD. Future work to focus on comparisons with chest pain patients with CVD will be useful to strengthen the use of miRNAs in early disease detection. It will also be interesting to understand if these markers are affected in cases of heart failure as these patients can present with chest pain and elevated troponins as well. Heart failure is one of the major causes of morbidity and mortality, and additional diagnostic assays will better aid in disease control. For more detailed analysis on these markers, it will be useful to understand their dynamics during disease manifestations. Prior studies had established in certain patient groups that some of the markers have appeared faster in blood than troponins (7). Understanding the dynamics in our patient group will allow better clinical management. Nonetheless, our study adds to the tools available for better management of the disease and showed that various cardiac related miRNAs is suitable and sensitive to changes in MI that can be used for diagnostics as well as for monitoring disease development in patients.
Conclusions
The present work examines the levels of miRNA in blood samples of NSTEMI patients suffering from AMI to establish their possible roles as a good disease biomarker. Our work focused on miR-1, miR-133, miR-208 and miR-499, given their close affinities to cardiac functions. Reference measurements showed clear distinct diagnostic performances comparable to troponin. We observed significant variability in a number of miRNAs tested and this could also possible link to post MI complications. Overall, miRNA measurements provide a fast and sensitive tool to complement existing techniques that can be beneficial to better manage the disease.
Acknowledgements
This work was supported by a research grant provided by the First Affiliated Hospital of Lanzhou University.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The study was approved by the institutional review board (No. 2543/00135) and written informed consent was obtained from all patients.
References
- Yusuf S, Reddy S, Ôunpuu S, et al. Global burden of cardiovascular diseases. Circulation 2001;104:2855-64. [Crossref] [PubMed]
- Abegunde DO, Mathers CD, Adam T, et al. The burden and costs of chronic diseases in low-income and middle-income countries. Lancet 2007;370:1929-38. [Crossref] [PubMed]
- Chen WW, Gao RL, Liu LS, et al. China cardiovascular diseases report 2015: a summary. J Geriatr Cardiol 2017;14:1-10. [PubMed]
- Adams JE, Schechtman KB, Landt Y, et al. Comparable detection of acute myocardial infarction by creatine kinase MB isoenzyme and cardiac troponin I. Clin Chem 1994;40:1291-5. [PubMed]
- Ravkilde J, Nissen H, Hørder M, et al. Independent prognostic value of serum creatine kinase isoenzyme MB mass, cardiac troponin T and myosin light chain levels in suspected acute myocardial infarction: analysis of 28 months of follow-up in 196 patients. J Am Coll Cardiol 1995;25:574-81. [Crossref] [PubMed]
- Goch A, Misiewicz P, Rysz J, et al. The clinical manifestation of myocardial infarction in elderly patients. Clin Cardiol 2009;32:E46-51. [Crossref] [PubMed]
- Chen C, Xu J, Huang F. Recent players in the field of acute myocardial infarction biomarkers: circulating cell-free DNA or microRNAs? Int J Cardiol 2013;168:2956-7. [Crossref] [PubMed]
- Maegdefessel L. The emerging role of microRNAs in cardiovascular disease. J Intern Med 2014;276:633-44. [Crossref] [PubMed]
- Baccarelli AA, Byun HM. Platelet mitochondrial DNA methylation: a potential new marker of cardiovascular disease. Clin Epigenetics 2015;7:44. [Crossref] [PubMed]
- Jylhävä J, Lehtimäki T, Jula A, et al. Circulating cell-free DNA is associated with cardiometabolic risk factors: the Health 2000 Survey. Atherosclerosis 2014;233:268-71. [Crossref] [PubMed]
- Creemers EE, Tijsen AJ, Pinto YM. Circulating microRNAs: novel biomarkers and extracellular communicators in cardiovascular disease? Circ Res 2012;110:483-95. [Crossref] [PubMed]
- Ai J, Zhang R, Li Y, et al. Circulating microRNA-1 as a potential novel biomarker for acute myocardial infarction. Biochem Biophys Res Commun 2010;391:73-7. [Crossref] [PubMed]
- Kuwabara Y, Ono K, Horie T, et al. Increased microRNA-1 and microRNA-133a levels in serum of patients with cardiovascular disease indicate the existence of myocardial damage. Circ Cardiovasc Genet 2011;4:446-54. [Crossref] [PubMed]
- Quiat D, Olson EN. MicroRNAs in cardiovascular disease: from pathogenesis to prevention and treatment. J Clin Invest 2013;123:11-8. [Crossref] [PubMed]
- Rana TM. Illuminating the silence: understanding the structure and function of small RNAs. Nat Rev Mol Cell Biol 2007;8:23-36. [Crossref] [PubMed]
- Johnson CD, Esquela-Kerscher A, Stefani G, et al. The let-7 microRNA represses cell proliferation pathways in human cells. Cancer Res 2007;67:7713-22. [Crossref] [PubMed]
- Frankel LB, Christoffersen NR, Jacobsen A, et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J Biol Chem 2008;283:1026-33. [Crossref] [PubMed]
- Yang B, Lin H, Xiao J, et al. The muscle-specific microRNA miR-1 regulates cardiac arrhythmogenic potential by targeting GJA1 and KCNJ2. Nat Med 2007;13:486-91. [Crossref] [PubMed]
- Townley-Tilson WHD, Callis TE, Wang D. MicroRNAs 1, 133, and 206: critical factors of skeletal and cardiac muscle development, function, and disease. Int J Biochem Cell Biol 2010;42:1252-5. [Crossref] [PubMed]
- Care A, Catalucci D, Felicetti F, et al. MicroRNA-133 controls cardiac hypertrophy. Nat Med 2007;13:613-8. [Crossref] [PubMed]
- Amsterdam EA, Wenger NK, Brindis RG, et al. 2014 AHA/ACC guideline for the management of patients with non-ST-elevation acute coronary syndromes: a report of the American College of Cardiology/American Heart Association Task Force on Practice Guidelines. Circulation 2014;130:e344-426. [Crossref] [PubMed]
- Challa PK, Smith KM, Conti CR. Initial presenting electrocardiogram as determinant for hospital admission in patients presenting to the emergency department with chest pain: a pilot investigation. Clin Cardiol 2007;30:558-61. [Crossref] [PubMed]
- Friesinger GC, Smith RF. Old age, left bundle branch block and acute myocardial infarction: a vexing and lethal combination. J Am Coll Cardiol 2000;36:713-6. [Crossref] [PubMed]
- Collinson PO, Stubbs PJ. Are troponins confusing? Heart 2003;89:1285-7. [Crossref] [PubMed]
- Rüegger S, Großhans H. MicroRNA turnover: when, how, and why. Trends Biochem Sci 2012;37:436-46. [Crossref] [PubMed]
- Christ M, Popp S, Pohlmann H, et al. Implementation of high sensitivity cardiac troponin T measurement in the emergency department. Am J Med 2010;123:1134-42. [Crossref] [PubMed]
- Twerenbold R, Reichlin T, Reiter M, et al. High-sensitive cardiac troponin: friend or foe. Swiss Med Wkly 2011;141. [PubMed]
- Filusch A, Giannitsis E, Katus HA, et al. High-sensitive troponin T: a novel biomarker for prognosis and disease severity in patients with pulmonary arterial hypertension. Clin Sci 2010;119:207-13. [Crossref] [PubMed]
- Long G, Wang F, Duan Q, et al. Human circulating microRNA-1 and microRNA-126 as potential novel indicators for acute myocardial infarction. Int J Biol Sci 2012;8:811. [Crossref] [PubMed]
- Olivieri F, Antonicelli R, Lorenzi M, et al. Diagnostic potential of circulating miR-499-5p in elderly patients with acute non ST-elevation myocardial infarction. Int J Cardiol 2013;167:531-6. [Crossref] [PubMed]
- Zhao Y, Ransom JF, Li A, et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1-2. Cell 2007;129:303-17. [Crossref] [PubMed]
- van Rooij E, Sutherland LB, Qi X, et al. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 2007;316:575-9. [Crossref] [PubMed]