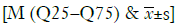The role of plasma miRNAs in the diagnosis of pulmonary nodules
Introduction
Based on new cancer data from China, lung cancer has the highest morbidity and mortality (1). Lung cancer is also the most prevalent cause of cancer-related death in the world (2). Although many lung cancer risk factors have been confirmed, such as age (3) and smoking history (4), among others, the prognosis of lung cancer is still poor, and the 5-year overall survival is approximately 15% (5). According to previous research, the 5-year overall survival of advanced lung cancer is approximately 6%. However, it is approximately 82% for early-stage lung cancer. Furthermore, only 10~15% of new lung cancer patients have clinical early-stage lung cancer. Therefore, the early diagnosis of lung cancer is important for improving prognosis. Additionally, it is important to find new methods to help improve the early diagnostic rate of lung cancer.
In recent years, liquid biopsy has emerged as a new detection technique, that can be applied to the diagnosis and monitoring of tumors (6). Moreover, microRNAs (miRNAs) are an important component of this technique. miRNAs are small, single-stranded, endogenous non-coding RNAs (7). More than 6,000 miRNAs have been discovered, and approximately half of them are human-related (8). They can promote or prevent tumor progression and metastasis, such as the function of proto-oncogenes and tumor suppressor genes. In recent years, some studies have shown that the expression of some miRNAs, such as miRNA-21 and miRNA-155 is upregulated in tumors. However, some miRNAs exhibit the opposite pattern, such as miRNA-34 and miRNA-145. This situation implies that it is helpful for us to detect the miRNA content to estimate the nature of the pulmonary nodules. At present, the tumor markers for lung cancer, which are used clinically, are carcinoembryonic antigen (CEA), cytokeratio-19 fragments (Cyfra21-1), neuron-specific enolase (NSE), squamous cell carcinoma antigen (SCCA) and so on. These tumor markers play a certain role in the diagnosis of lung cancer, but their specificity and sensitivity are not enough. Furthermore, these tumor markers are not elevated in many early lung cancer patients. Therefore, it is imperative to find new molecular biomarkers to help diagnose early-stage lung cancer. Additionally, miRNA can steadily exist in human plasma, express specifically in some tumors, and are not degraded easily by RNA enzymes, which make miRNA a potential new molecular biomarker (9).
After the first miRNA (miRNA let-7) was found, more than 300 miRNAs have been identified and considered to be related to human tumors. The role of miRNAs in the diagnosis of lung cancer has been only partly explored previously. Some studies have revealed that miRNAs play an important role in the development, progression, diagnosis, treatment, and prognosis of lung cancer. Following a lot of previous studies which have demonstrated the value of miRNAs for diagnosing pulmonary nodules (10-13), we measured 12 miRNAs (miRNA-17, -146a, -200b, -182, -155, -221, -205, -126, -7, -21, -145, and miRNA-210) in this study. The purpose of this study was to investigate the difference in the expression of miRNAs between benign pulmonary nodules and malignant pulmonary nodules and explored the role of plasma miRNAs in the diagnosis of pulmonary nodules.
Methods
Patient selection
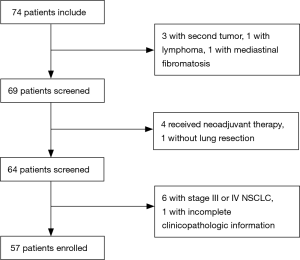
The study was approved by the Ethics Committee of Sun Yat-sen University Cancer Center (No. B2017-050) and written informed consent was obtained from all patients. In this retrospective study, we collected people who were diagnosed with a pulmonary nodule by thoracic computed tomography (CT) and were scheduled to receive surgery from July 2016 to July 2017 at Sun Yat-sen University Cancer Center. In this study, patients were enrolled following the eligibility criteria: (I) all of the patients were diagnosed with a pulmonary nodule by CT before the surgery; (II) all of the patients had pathologically confirmed stage 0–II non-small cell lung cancer (NSCLC) or benign lung disease according to the International Association for the Study of Lung Cancer (7th version); and (III) all of the patients had complete clinicopathologic information. The exclusion criteria in this study were as follows: (I) patients received neoadjuvant therapy, including radiotherapy, chemotherapy, or chemoradiotherapy; (II) patients had a second primary tumor; (III) patients were not pathologically confirmed after surgery; and (IV) patients had stage III or IV lung cancer. Finally, 57 patients were included in our study. The flow was shown in Figure 1.
Detection procedure
We drew approximately 5 mL of venous blood from all patients before the operation. We spun these samples in a centrifuge and obtained plasma. Then, the plasma was immediately stored at −80 °C. The MiRNeasy Serum/Plasma Kit (Qiagen, Germany) was used to extract the total RNA from the plasma (200 µL). The Omniscript RT Kit (Qiagen, Germany) was used to accomplish reverse transcription (RT) reactions. The levers of miRNAs were detected by real-time quantitative polymerase chain reaction (RT-PCR) using the SYBR Green Mix (Qiagen, Cat No. 208054). The relative miRNA expression was calculated using the 2−△△Ct method. In order to eliminate the bias during sample storage or RNA isolation, the cel-miR-39 was chosen as the inner control.
Statistical analysis
Difference in the plasma miRNA levels between NSCLC group and benign pulmonary nodules group was evaluated by Mann-Whitney U test and t-test. T-test, analysis of variance, Mann-Whitney U test, and Kruskal-Wallis test were used to evaluate the relationship between the expression levels of the plasma miRNAs and clinicopathologic characteristics. We used the receiver operating characteristic (ROC) curve to assess the efficacy of plasma miRNAs in the diagnosis of NSCLC. We used the Shapiro-Wilk test for the test for normal distribution. When the data was not normally distributed, we used the Wilcoxon rank-sum test to analyze data. T test was used for univariate analysis when the data was normally distributed. However, for multiple sets of measurement data, using analysis of variance. Results were Log10 transformed for analysis because of the magnitude of miRNA levels measured. Statistical analyses were performed using SPSS software version 21 (SPSS Inc., Chicago, IL). P values <0.05 was considered statistically significant.
Results
Patient characteristics
Fifty-seven patients were enrolled in this study, including 15 benign pulmonary nodules patients and 42 malignant pulmonary nodules patients; 26 men and 31 women; 32 patients age ≤60 and 25 patients age >60. Their median age was 59 years (range from 43 to 81 years). Among the 42 NSCLC patients, 20 men and 22 women were included. This included twenty-one patients age ≤60 and 21 patients age >60. The age of the NSCLC patients ranged from 43–77 years (median 60.5 years). Other characteristics are shown in Tables 1 and 2.

Full table
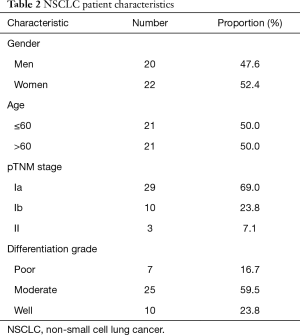
Full table
Association of the expression of miRNAs with clinical characteristics
As is shown in Table 3, we used the T test, Mann-Whitney U test, analysis of variance, and Kruskal-Wallis test to analyze the connection between the expression of the miRNAs and the clinical characteristics in 42 NSCLC patients. The results show that the miRNA expression (miRNA-17, -146a, -200b, -182, -155, -221, -205, -126, -7, -21, -145, -210) in NSCLC was not associated with gender, age, pTNM stage and differentiation grade.
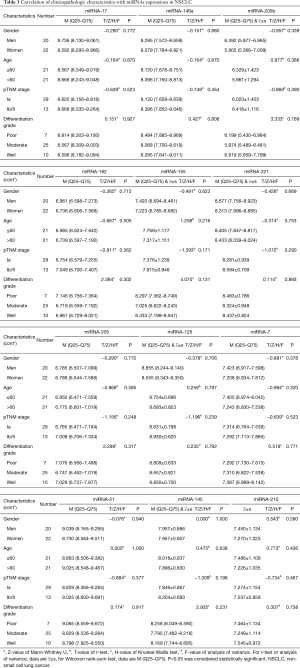
Full table
The expression of 12 miRNAs
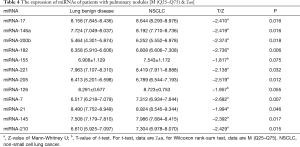
Fifty-seven patients were divided into two groups according to the characteristics of their pulmonary nodule (benign or malignant). The t-test and Mann-Whitney U test were used to compare the differences in the expression of the miRNAs of the two groups (NSCLC group and benign pulmonary nodules group). The results are listed in Table 4, which indicate that the expression levels of plasma miRNA-17, -146a, -200b, -182, -221, -205, -7, -21, -145, and miRNA-210 in NSCLC patients are significantly higher than those in the benign pulmonary nodules patients (P<0.05). However, for miRNA-155 and miRNA-126, there was not a significant difference between these two groups, the P values are 0.075 and 0.055, respectively.
The role of plasma miRNAs in the diagnosis of pulmonary nodules
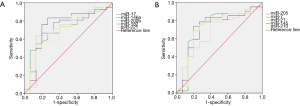
We evaluated the value of the miRNA spectrum in the diagnosis of pulmonary nodules using the ROC curve. As shown in Table 5, the sensitivity and specificity were 66.7% and 80.0% for miRNA-17, 54.8% and 86.7% for miRNA-146a, 64.3% and 86.7% for miRNA-200b, 83.3% and 73.3% for miRNA-182, 54.8% and 80.0% for miRNA-221, 73.8% and 80.0% for miRNA-205, 78.6% and 73.3% for miRNA-7, 78.6% and 60.0% for miRNA-21, 78.6% and 73.3% for miRNA-145, 76.2% and 73.3% for miRNA-210, respectively. The ROC curve of the 10 miRNAs is show in Figure 2. To sum up, the plasma miRNA-17, -146a, -200b, -182, -221, -205, -7, -21, -145, and miRNA-210 have a relatively high sensitivity (54.8–83.3%) and specificity (60.0–86.7%) for the diagnosis of NSCLC.

Full table
Discussion
Lung cancer is the most common cause of cancer-related death in the world (14). The prognosis of lung cancer is still poor despite the advances in cancer treatments over the past decades. Late diagnosis of NSCLC is one of the most important reasons. Hence, new methods are urgently needed to diagnose early lung cancer. Previous studies indicate that miRNAs may be the potential diagnostic biomarkers of lung cancer (13,15).
In this study, our results revealed that the expression levels of miRNAs in NSCLC patients were not related to other clinical and pathologic features. The available literature shows different conclusions in this respect. For example, Zhang et al. identified that the expression level of miRNA-221 was not relevant to the TNM stage (16). However, Yin et al. found that the expression level of miRNA-221 was correlated with the TNM stage (17). The relationship between the clinicopathological characteristics of NSCLC patients and the expression of miRNAs need be explored further.
We discovered that there were significant differences in the expression of the miRNAs between NSCLC patients and benign pulmonary nodules patients (miRNA-17, -146a, -200b, -182, -221, -205, -7, -21, -145 and miRNA-210 except miRNA-155, miRNA-126). The expressions of miRNA-17 (18), -182 (19,20), -221 (16,17), -21 (21,22), -210 (23) were significantly higher in NSCLC patients compared with that of benign pulmonary nodules, which confirmed the previous research. The expression of miRNA-146a was increased and downregulated in a variety of carcinomas (24), while miRNA-146a expression was increased in NSCLC patients in our study compared with that of nonmalignant disease. Han et al. demonstrated that miRNA-200b expression was upregulated in bladder urothelial carcinoma (25). However, it was downregulated in liver cancer and renal cell carcinoma (26,27). We found that it was upregulated in NSCLC. The research concerning the expression level of miRNA-200b in lung cancer was inadequate and it was upregulated in NSCLC in our study. Xiong et al. reported that miRNA-7 was downregulated in NSCLC (28), while Ye et al. revealed that miRNA-145 was also downregulated in NSCLC (29). These results and our conclusions were inconsistent. More samples are needed to verify this phenomenon in the future. While for miRNA-155 and miRNA-126, there was not a significant difference between NSCLC group and benign pulmonary nodules group, the P values are 0.075 and 0.055, respectively. Their P values are closed to the 0.05, especially miRNA-126 (P=0.055). Maybe due to the small sample size.
We used ROC curves to determine the diagnostic value of 10 miRNAs (miRNA-17, -146a, -200b, -182, -221, -205, -7, -21, -145, -210) for lung nodules. A number of studies have determined that miRNAs may be potential molecular biomarkers for early carcinoma including lung cancer (15,30). Zhang et al. demonstrated that plasma miRNA-145 and miRNA-21 can be used as potential biomarkers for diagnosing lung cancer. The sensitivity and specificity of miRNA-145 were 80.6% and 89.2%, and 77.5% and 85.5% for miRNA-21, respectively (31). Sromek et al. found that the combination of miRNA-9, -16, -205, and -486 can serve as potential NSCLC biomarkers with 80% sensitivity and 95% specificity (32). In the present study, the miRNAs have a relatively high sensitivity (54.8–83.3%) and specificity (60.0–86.7%) for the diagnosis of NSCLC. Currently, growing numbers of lung nodule patients have been discovered because of the increasingly widespread use of low-dose spiral computed tomography (LDSCT) and CT. However, the specificity of LDSCT/CT is limited, and false positive rate is relatively high (33). Additionally, our results reveal that, with the help of LDSCT/CT, miRNAs can provide an auxiliary diagnosis of lung nodules. Based on previous literatures, cancer-associated miRNAs played crucial roles in tumorigenesis and may be as diagnostic and prognostic biomarkers in cancer, including carcinoma of lungs (9,34). As a new latent molecular biomarker, miRNA has its advantages: non-invasion, easy sample management, cost-effectiveness, which make it may influence the future diagnostic method of lung cancer. This study focused on 12 miRNAs and selected 10 miRNAs which contribute to produce the prediction in distinguishing lung carcinoma patients from benign subjects according to the result. This pilot study provided the theory foundation for the development of the research—the role of plasma miRNAs in early diagnosis of lung cancer.
This study has several limitations, our data was collected retrospectively and a single team. This approach may lead to a selection bias. Furthermore, was the small sample size, which may be the reason for no difference between the NSCLC group and benign pulmonary nodules group for the levels of miRNA-155 and miRNA-126. Designing a prospective study with a large sample size in the future, conducting combined diagnosis of miRNAs and constructing a risk score model would be more valuable.
In summary, 10 miRNAs (miRNA-17, -146a, -200b, -182, -221, -205, -7, -21, -145, -210) were found that they had relatively high diagnostic efficacy to discriminate the nature of pulmonary nodules. Understanding miRNAs have the potential to become novel non-invasive biomarkers for the diagnosis of early-stage lung cancer, and as the complementary of LDSCT screening procedure. MiRNAs may contribute to the development of diagnostic techniques for early-stage lung cancer in the future.
Acknowledgements
Funding: This study was funded by National Key Research and Development Program, China (No. 2016YFC0905400).
Footnote
Conflicts of Interest: The authors have no conflict of interest to declare.
Ethical Statement: The study was approved by the Ethics Committee of Sun Yat-sen University Cancer Center (No. B2017-050) and written informed consent was obtained from all patients.
References
- Chen W, Zheng R, Baade PD, et al. Cancer statistics in China, 2015. CA Cancer J Clin 2016;66:115-32. [Crossref] [PubMed]
- Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87-108. [Crossref] [PubMed]
- Salmeron D, Chirlaque MD, Isabel Izarzugaza M, et al. Lung cancer prognosis in Spain: the role of histology, age and sex. Respir Med 2012;106:1301-8. [Crossref] [PubMed]
- Kogure Y, Ando M, Saka H, et al. Histology and smoking status predict survival of patients with advanced non-small-cell lung cancer. Results of West Japan Oncology Group (WJOG) Study 3906L. J Thorac Oncol 2013;8:753-8. [Crossref] [PubMed]
- Wu YL, Zhou Q. Clinical trials and biomarker research on lung cancer in China. Expert Opin Ther Targets 2012;16 Suppl 1:S45-50. [Crossref] [PubMed]
- Crowley E, Di Nicolantonio F, Loupakis F, et al. Liquid biopsy: monitoring cancer-genetics in the blood. Nat Rev Clin Oncol 2013;10:472-84. [Crossref] [PubMed]
- Ambros V. The functions of animal microRNAs. Nature 2004;431:350-5. [Crossref] [PubMed]
- Wang Y, Lee CG. MicroRNA and cancer--focus on apoptosis. J Cell Mol Med 2009;13:12-23. [Crossref] [PubMed]
- Ferracin M, Veronese A, Negrini M. Micromarkers: miRNAs in cancer diagnosis and prognosis. Expert Rev Mol Diagn 2010;10:297-308. [Crossref] [PubMed]
- Zheng D, Haddadin S, Wang Y, et al. Plasma microRNAs as novel biomarkers for early detection of lung cancer. Int J Clin Exp Pathol 2011;4:575-86. [PubMed]
- Shen J, Todd NW, Zhang H, et al. Plasma microRNAs as potential biomarkers for non-small-cell lung cancer. Lab Invest 2011;91:579-87. [Crossref] [PubMed]
- Chen X, Ba Y, Ma L, et al. Characterization of microRNAs in serum: a novel class of biomarkers for diagnosis of cancer and other diseases. Cell Res 2008;18:997-1006. [Crossref] [PubMed]
- Wang RJ, Zheng YH, Wang P, et al. Serum miR-125a-5p, miR-145 and miR-146a as diagnostic biomarkers in non-small cell lung cancer. Int J Clin Exp Pathol 2015;8:765-71. [PubMed]
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2015. CA Cancer J Clin 2015;65:5-29. [Crossref] [PubMed]
- Wozniak MB, Scelo G, Muller DC, et al. Circulating MicroRNAs as Non-Invasive Biomarkers for Early Detection of Non-Small-Cell Lung Cancer. PLoS One 2015;10. [Crossref] [PubMed]
- Zhang Y, Zhao Y, Sun S, et al. Overexpression of MicroRNA-221 is associated with poor prognosis in non-small cell lung cancer patients. Tumour Biol 2016;37:10155-60. [Crossref] [PubMed]
- Yin Z, Xu M, Li P. miRNA-221 acts as an oncogenic role by directly targeting TIMP2 in non-small-cell lung carcinoma. Gene 2017;620:46-53. [Crossref] [PubMed]
- Chen Q, Si Q, Xiao S, et al. Prognostic significance of serum miR-17-5p in lung cancer. Med Oncol 2013;30:353. [Crossref] [PubMed]
- Wang M, Wang Y, Zang W, et al. Downregulation of microRNA-182 inhibits cell growth and invasion by targeting programmed cell death 4 in human lung adenocarcinoma cells. Tumour Biol 2014;35:39-46. [Crossref] [PubMed]
- Saus E, Soria V, Escaramis G, et al. Genetic variants and abnormal processing of pre-miR-182, a circadian clock modulator, in major depression patients with late insomnia. Hum Mol Genet 2010;19:4017-25. [Crossref] [PubMed]
- Li X, Zhang Q, Jin X, et al. Combining biomarker, imaging, and clinical features to distinguish pulmonary nodules. World J Surg Oncol 2017;15:107. [Crossref] [PubMed]
- Capodanno A, Boldrini L, Proietti A, et al. Let-7g and miR-21 expression in non-small cell lung cancer: correlation with clinicopathological and molecular features. Int J Oncol 2013;43:765-74. [Crossref] [PubMed]
- Raponi M, Dossey L, Jatkoe T, et al. MicroRNA classifiers for predicting prognosis of squamous cell lung cancer. Cancer Res 2009;69:5776-83. [Crossref] [PubMed]
- Labbaye C, Testa U. The emerging role of MIR-146A in the control of hematopoiesis, immune function and cancer. J Hematol Oncol 2012;5:13. [Crossref] [PubMed]
- Han Y, Chen J, Zhao X, et al. MicroRNA expression signatures of bladder cancer revealed by deep sequencing. PLoS One 2011;6. [Crossref] [PubMed]
- Murakami Y, Toyoda H, Tanaka M, et al. The progression of liver fibrosis is related with overexpression of the miR-199 and 200 families. PLoS One 2011;6. [Crossref] [PubMed]
- Slaby O, Jancovicova J, Lakomy R, et al. Expression of miRNA-106b in conventional renal cell carcinoma is a potential marker for prediction of early metastasis after nephrectomy. J Exp Clin Cancer Res 2010;29:90. [Crossref] [PubMed]
- Xiong S, Zheng Y, Jiang P, et al. MicroRNA-7 inhibits the growth of human non-small cell lung cancer A549 cells through targeting BCL-2. Int J Biol Sci 2011;7:805-14. [Crossref] [PubMed]
- Ye Z, Shen N, Weng Y, et al. Low miR-145 silenced by DNA methylation promotes NSCLC cell proliferation, migration and invasion by targeting mucin 1. Cancer Biol Ther 2015;16:1071-9. [Crossref] [PubMed]
- Cho WC. MicroRNAs: potential biomarkers for cancer diagnosis, prognosis and targets for therapy. Int J Biochem Cell Biol 2010;42:1273-81. [Crossref] [PubMed]
- Zhang H, Mao F, Shen T, et al. Plasma miR-145, miR-20a, miR-21 and miR-223 as novel biomarkers for screening early-stage non-small cell lung cancer. Oncol Lett 2017;13:669-76. [Crossref] [PubMed]
- Sromek M, Glogowski M, Chechlinska M, et al. Changes in plasma miR-9, miR-16, miR-205 and miR-486 levels after non-small cell lung cancer resection. Cell Oncol (Dordr) 2017;40:529-36. [Crossref] [PubMed]
- Swensen SJ, Jett JR, Sloan JA, et al. Screening for lung cancer with low-dose spiral computed tomography. Am J Respir Crit Care Med 2002;165:508-13. [Crossref] [PubMed]
- Wang QZ, Xu W, Habib N, et al. Potential uses of microRNA in lung cancer diagnosis, prognosis, and therapy. Curr Cancer Drug Targets 2009;9:572-94. [Crossref] [PubMed]