
Wide phenotypic variability in RSPH9-associated primary ciliary dyskinesia: review of a case-series from Cyprus
Introduction
Primary ciliary dyskinesia (PCD; MIM 244400) is an inherited ciliary motility disorder characterized by chronic lung disease, randomized body laterality and infertility. Chronic lung disease in PCD is the result of impaired mucociliary clearance which typically leads to recurrent airway infections, bronchiectasis, and progressive loss of lung function (1). Diagnosis is often delayed, especially when typical clinical signs, such as situs inversus, are missing (2). The cited incidence of 1:10,000–20,000 for PCD, may be an underestimation of the true incidence due to the phenotypic heterogeneity of the disease and the difficulty in performing and interpreting diagnostic tests (2). The phenotypic heterogeneity is also expressed in key diagnostic features and as a consequence there is no “gold standard” diagnostic test of choice suitable in all cases (3), but a combination of tests which primarily includes nasal nitric oxide (nNO) (4), high speed video microscopy (HSVM) (5) and transmission electron microscopy (TEM) (6).
The axonemes of motile cilia are composed of more than 200 proteins (7) that form a characteristic structure of 9 peripheral microtubule doublets each bearing a pair of outer (ODA) and inner dynein arms (IDA), a central microtubule pair, and 9 connecting radial spokes (8). To date mutations in at least 40 genes have been reported to cause PCD, accounting for up to 75% of the genetic diagnoses in various PCD cohorts worldwide (9,10). Defects in radial spoke head proteins RSPH1, 3, 4a and 9, have been linked with PCD (11-14).
In 2009, Castleman et al., demonstrated for the first time, that the mutation p.Lys268del in the RSPH9 gene in three children of Bedouin origin from Israel and the United Arabic Emirates causes an unusual intermittent loss of the central pair, presenting in a small proportion of cilia cross sections with 9+0 structure and abnormal circular movement, but with ciliary beat frequency within the normal range (14). The p.Lys268del mutation in RSPH9 gene was subsequently reported in three siblings with PCD from southwest Saudi Arabia, suggesting that this genetic alteration is geographically widespread in the Arabian Peninsula (13). The largest cohort of PCD patients with isolated RSPH9 gene mutations was reported by Frommer et al. in 2015, who described eight patients of Israeli and North Western European origins, of whom five with bi-allelic homozygous loss-of-function mutations [(c.466C>T, p.(Arg156*); c.2T>C, p.(Met1?); c.523–1G>C, [?] and c.610A>T, p.(Lys204*) and c.2T>C, p.(Met1?) and c.610A>T, p.(Lys204*)], two with homozygous in-frame deletions (c.801_803delGAA, p.(Lys268del) and one with a homozygous missense mutation (c.752A>G, p.(His251Arg) (11). There is no information on the prevalence of RSPH9 gene mutations in other populations, especially in neighboring countries in the Eastern Mediterranean basin.
All previous studies on RSPH9-associated PCD, aimed to identify the role of RSPH9 gene mutations in the causation of PCD and thus followed a linkage analysis approach in patients who presented with classic PCD clinical features (11,13,14). However, RSPH9 gene mutations encode intermittent ultrastructural defects, which are difficult to diagnose by TEM, and subtle ciliary beating abnormalities on HSVM. Thus, it is very likely that many patients are not diagnosed and, as a result, the true clinical spectrum of RSPH9 gene mutations remains largely unknown.
The aim of this study was to perform whole exome sequencing in two unrelated probands of suspect patients from Cyprus with aberrant TEM and HSVM findings, similar to those described previously in PCD patients bearing pathogenic RSPH9 mutations and review the diagnostic and clinical phenotype in the confirmed cases.
Methods
Diagnostic testing
All subjects had standardized PCD diagnostic testing which included nNO measurements and HSVM and TEM studies. nNO levels were measured by a chemiluminescence analyzer (CLD88sp, Ecomedics Switzerland) (15). Cilia studies were performed directly in samples of respiratory epithelial cells obtained with brushing the inferior nasal turbinate in all patients. Nasal brushings were carried out in the absence of any respiratory infection for at least 4 weeks. A ZEISS Axiophot microscope (ZEISS, Germany) equipped with a Basler scA640 firewire video camera (Basler Vision Technologies, Germany) was used for the evaluation of ciliary beat pattern (CBP) and Sisson-Ammons Video Analysis System (SAVA system) (16) was used for the quantification of ciliary beat frequency (CBF) at 37 °C. TEM analysis was carried out as described previously (17). HSVM and TEM findings were evaluated by two independent reviewers and final appraisal of the findings was performed by a multidisciplinary team of experts.
Genetic testing
All patients or their guardians gave written informed consent for molecular testing and participation in the study. DNA was extracted from peripheral blood and whole exome sequencing (WES) was performed using the Nextera Rapid Capture exome kit (Illumina) on a NextSeq500 (Illumina) next generation sequencing platform. Paired-end reads of 101 bp were generated with mean coverage sequencing depth of 100×. A previously established bioinformatics pipeline was used to process and analyze WES data. Specifically, Burrows-Wheeler Aligner (BWA) was used to map and align paired-end reads to the human reference genome (version GRCh37/hg19). PCR duplicates as well as non-uniquely mapped reads were filtered out to ensure accurate alignment and variant calling rates. Variant calling was performed using GATK’s Haplotype Caller. More information on the established bioinformatics pipeline can be found elsewhere (18). The identified RSPH9 mutation was confirmed by Sanger sequencing in all seven patients and tested for disease segregation in familial DNA samples. The frequency of the detected variant was also determined by Sanger sequencing in a pool of 400 unrelated Cypriot healthy individuals (based on a health status questionnaire), which included 150 samples from the same district as the proband cases.
Clinical phenotype
Basic demographic data including age at diagnosis and current age (as of middle of 2017) were recorded. Clinical history information was obtained by a standardized questionnaire addressed to patients or guardians, which covered important manifestations i.e., laterality defects, neonatal respiratory distress, chronic rhinorrhea and or rhinosinusitis, cough, wheezing, pneumonia [report of any episode(s) of pneumonia ever], nasal polyps, report of lobectomy, grommets insertion, sinus surgery, hearing problems, fertility difficulties (where appropriate) and chest computed tomography (CT) confirmed bronchiectasis. We also recorded spirometric indices at presentation. Forced expiratory volume in one sec (FEV1) and forced vital capacity (FVC) were expressed as z-scores of the predicted for the patient’s height, age and gender (19). Body mass index (BMI) (kg/m2) was expressed as age- and gender-specific z-scores based on the United States’ Centers for Disease Control 2000 growth charts (20).
Statistical analysis
Categorical variables are presented as frequencies (%) and continuous variables are expressed as median (range). Summary statistics were calculated using STATA 12 (Version 12, StataCorp, College Station, TX, USA).
Results
Presentation of the probands
Family 1
Index case 1, who led to the identification of suspect patients in the first family, female aged 6.6 years, presented with situs solitus (Table 1), and typical PCD manifestations including chronic rhinitis, chronic wet cough since birth and a history of past pneumonia, neonatal respiratory distress and grommets insertion (Table 2). nNO production was abnormally low at 41.7 nL/min. Nasal brush sampling for HSVM revealed a CBF of 9.35 Hz and rotational CBP. On TEM there were abnormalities of the central apparatus in 40% of the cross sections with no abnormality in ODA and IDA (Table 1). Nasal brushing was repeated two more times and the findings on TEM and HSVM were the same as those of the first brushing. Two siblings of the index case (cases 2 & 3) (See pedigree of family 1 in the online supplement section of the manuscript, Figure S1) had situs solitus, chronic rhinitis and recurrent wet cough since school age, but no other suspect manifestations for PCD (Table 1). In the following 3 years, the two siblings were invited at the age of 14.6 and 16.9 years for PCD diagnostic work-up, which was repeated three times in each case to confirm findings.
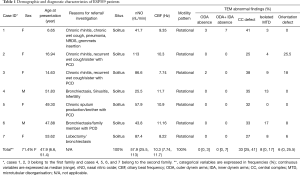
Full table
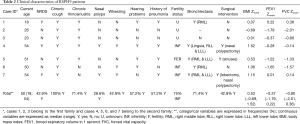
Full table
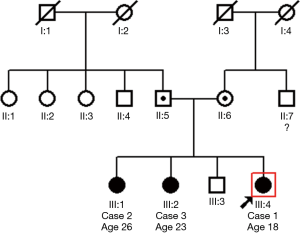
Family 2
Index case 4 from the second family, male aged 51.8 years, displayed situs solitus, and typical PCD manifestations including chronic rhino-sinusitis, chronic wet cough since school age and a history of typical PCD manifestations i.e., recurrent pneumonias, bronchiectasis, nasal polypectomy, infertility, neonatal respiratory distress, and reduced hearing (Table 2). nNO production was abnormally low at 25.5 nL/min. HSVM revealed a CBF of 11.7 Hz and a rotational CBP. On TEM there were abnormalities of the central apparatus in 35% of the cross sections (Table 1). The nasal cilia biopsy was repeated and TEM and HSVM findings were consistent. In the family of the index case there were two siblings (cases 5 & 6) and one first cousin (case 7) (See pedigree of family 2 in the online supplement section of the manuscript, Figure S2) with situs solitus and variable but milder chronic upper and lower respiratory system manifestations (Table 2). In the following year, the three suspect patients were invited at the ages of 49.3, 47.9 and 53.6 years for PCD diagnostic work-up, which gave similar findings to the index case.
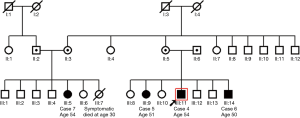
Genetic results
The two index cases were homozygous for the RSPH9 splice site mutation c.670+2T>C in intron 4 which can be classified as pathogenic based on the latest Association for Clinical Genomic Science (ACGS) Best Practice Guidelines for Variant Classification (21). The index cases from both families were analyzed by exome sequencing, hence all known PCD genes were sequenced and mutations in any other PCD-associated genes apart from RSPH9 were not detected. In addition, all family members tested (n=7), who had similar aberrant TEM and HSVM diagnostic features, were also homozygous for this splice site mutation. This mutation has not been reported before in the literature. In silico analysis of the mutant sequence using three different splicing prediction software (Human Splicing Finder, NetGene2 and NNSplice) concluded that this mutation causes an alteration of the wild type donor site, most probably affecting splicing and thus leading to truncation of the protein. Only one out of the 400 healthy controls tested was heterozygous for the RSPH9 c.670+2T>C mutation. It is of interest that this individual originates from the same district as the two index cases.
Diagnostic and clinical phenotype in the confirmed cases
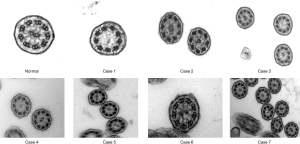
We investigated 7 individuals (5 females) from the two families with age range at presentation 6.6 to 51.4 years (median 47.9 years) with no laterality defects (situs solitus). Cases 2, 3 and 6 presented with only mild manifestations, such as chronic nasal secretions, recurrent cough, normal spirometry and higher nasal nitric oxide. The average nNO production values ranged from 25.5 to 113 nL/min (median 57.9 nL/min) (Table 1). Three of the seven subjects had nNO levels higher than 77 nL/min, which is the disease-specific cutoff in PCD (19). On HSVM, the average CBF was 7.74 to 11.7 Hz (median 10.3 Hz), whereas rotational CBP was displayed on some of the examined wedges in all patients. Characteristic HSVM clips of the rotational beating pattern from all patients are presented in the online supplement section of the manuscript (Figures S3-S9). On TEM, there were abnormalities of the central apparatus in all cases which ranged from 25% to 41% (median 33%) of the examined cross sections. Characteristic TEM photos of the ciliary structural abnormalities from all patients are presented in the online supplement section of the manuscript (Figure S10). Isolated microtubular disorganization (3% to 17% of cross sections) was noted in 6 patients and orientation defects (6% to 25.5% of cross sections) were noted in 4 patients which were attributed to secondary changes. No significant ODA/IDA abnormalities were observed (Table 1).







All patients presented with a history of chronic or recurrent wet cough, but in 5 of them the symptoms reportedly started at primary school age. Out of the 7 patients, 3 (43%) reported neonatal respiratory distress, 5 (71%) had recurrent or chronic rhinosinusitis and 3 (43%) complained for wheezing. Bronchiectasis of widely varying severity and extent was documented in 5 patients (71%). Four patients had a history of pneumonia(s) and 4 experienced hearing problems (57%). Three patients had undergone surgical procedures such as polypectomy or sinuses surgery and lobectomy, and 3 of the 4 patients (75%) in child bearing age had fertility issues (Table 2).
BMI-z scores were above 0 in 6 of the 7 patients (median: 0.53; range: −0.69 to 1.52), whereas FEV1 (median: −0.37; range: −1.79 to 0.22) and FVC (median: −0.80; range: −2.01 to 0.36) were slightly lower but within the normal range (Table 2).
Discussion
We reviewed the first case-series of seven patients with RSPH9-associated PCD disease that is caused by the same, homozygous novel splice site mutation c.670+2T>C in intron 4. The patients come from two families that are reportedly non-consanguineous, even though they originate from the same district of South Cyprus. However, the rarity of the identified novel mutation, found in homozygosity in the two families, supports their common ancestry. The fact that molecular screening of 400 population-based healthy control samples from across the country for RSPH9 c.670+2T>C mutation revealed only one heterozygous carrier originating from the same district as the probands, also points towards a possible founder’s effect.
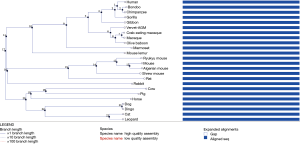
Although no RNA sample was available to examine the effect of the RSPH9 c.670+2T>C mutation on splicing, in silico analysis has demonstrated that it causes an abolishment of the existing donor site, resulting in a frameshift and a premature stop codon. The position of the mutation is conserved in the interspecies comparison with RSPH9 orthologues from 24 Eutherian mammals (Figure S11) and in silico assessment for the effect of the mutation on splicing was performed for both transcripts (NM_001193341.1 and NM_152732.4) of the RSPH9 gene. In both cases, in silico assessment reached the same conclusion (abolishment of the existing donor site resulting in protein truncation). This is a novel mutation that was identified for the first time in our patient case series hence it is absent from all gene variant databases. Furthermore, based on RNA-seq analysis, which was performed on human tissue samples in order to determine tissue-specificity of all protein-coding genes (22), there is a biased expression of RSPH9 in testis, lung and brain. Hence, protein truncation which may lead to reduced or absent RSPH9 expression in the lung can explain the clinical phenotype of our patients.
Previous reports on RSPH9-PCD described patients with the classical clinical features for PCD (11,13,14). Our study expands the clinical spectrum, that we consider to be PCD, since some cases (Cases 2, 3 and 6) presented with only mild manifestations, such as chronic nasal secretions, recurrent cough, normal spirometry and higher nasal nitric oxide. These cases would not normally have risen suspicion to undergo specialist diagnostic work-up for PCD, if they did not belong to families of patients with more typical disease manifestations. This is particularly important for genotypic alterations of RSPH9 since laterality defects have not been observed in these cases (14), thus permitting for important diagnostic uncertainty either in the early stages of respiratory disease or in cases with a milder clinical phenotype. Phenotypic consistency in our series was more profound as far as ultrastructural ciliary alterations and ciliary motility patterns are concerned. All patients had about the same percentage of ciliary cross sections with central pair defects on TEM and displayed a rotational beating pattern on HSVM. In contrast, beating frequency was near normal (>11 Hz) in two (23), as opposed to the rest of the patients. In terms of nNO production, levels were above the reported threshold of 77 nL/min for diagnosis of PCD (24) in three patients. The quoted prevalence of 1:10,000 to 1:20,000 for PCD is widely considered to be an underestimation of the true prevalence (2). The increasing discovery of new PCD genes and the wider adoption of genetic diagnostics are expected to reveal milder forms of the disease. A similar pattern was observed in the case of cystic fibrosis (CF) following the establishment of genetic diagnosis that signified the expansion of the disease clinical spectrum with the inclusion of many mild or atypical clinical phenotypes (25,26). Although it is estimated that the known genes account for up to 75% of the genetic diagnoses in various PCD cohorts worldwide (9,10), several recently discovered genes are encoding for subtle ciliary phenotypic abnormalities (27-31), which may change the clinical spectrum and the phenotypic severity of the disease in the next few years. Nevertheless, even in cases attributed to a specific defect in a PCD gene, the presence of genetic alterations in other genes involved in innate immunity (32) or exposure to adverse environmental factors such as environmental tobacco smoke (33) has been recently demonstrated to have modifying effects on clinical manifestations and induce variability in pulmonary function.
Our data suggest that in non-CF, non-Kartagener’s chronic respiratory disease, when there is difficulty to confirm PCD diagnosis with standard diagnostic tests, genetic testing must be undertaken, at least for the RSPH9 gene and perhaps other genes that encode for mild alterations in cilia ultrastructure and motility and cause atypical clinical manifestations. When definitive diagnosis is made, appropriate follow-up with regular surveillance of pulmonary function and respiratory microbiology as well as use of antibiotics targeted to pathogens, can prevent or minimize many of the disease consequences, which build up from childhood to adulthood. Similar approaches for expanding diagnostic screening in wider groups with respiratory disease for recovery of possible PCD patients have been proposed by previous reports on RSPH9-associated PCD disease (13).
The relatively small number of the presented cases is a limitation for our study. This is largely inherent in PCD studies due to the high number of the involved genes and the rarity of the cases attributed to a specific new mutation, which does not allow for statistical comparisons of the clinical phenotypic features. However, we feel that even the descriptive presentation of the diverse clinical characteristics relating to this new mutation has important implications. It is evident that further studies are required to elicit genotype-phenotype associations in PCD, as well as larger studies to assess the phenotypic variability in patients with the same genotypic alterations in PCD-associated genes. These studies are expected to be more complex than similar studies performed previously in CF (34,35) due to the many genes that are implicated in PCD causation. Nevertheless, the recent establishment of large international PCD registries (36,37) with several thousands of PCD patients offers the potential for such studies in the near future.
In conclusion, this case-series demonstrates the wide phenotypic variability of RSPH9-associated PCD disease, which in mild or atypical clinical presentations would not raise suspicion for specialist diagnostic work-up for PCD. We highlight the importance of wider adoption of genetic diagnostics in non-CF, non-Kartagener’s chronic respiratory disease, especially for genes that encode subtle ciliary phenotypic abnormalities, and help establish appropriate follow-up and introduction of careful management from early on life. Larger studies are needed on genotype-phenotype associations and phenotypic variability of alterations in PCD-associated genes.
Acknowledgments
The authors would also like to acknowledge the contribution of Dr. Nicos Middleton in English language editing of the manuscript.
Funding: The authors acknowledge funding from EU 7th Framework Program EC-GA No. 305404 BESTCILIA and Cyprus Research Promotion Foundation (IPE/KY/ROU-0311/11). PK Yiallouros, P Kouis and M Kalyva are members of EU-funded COST Action BEAT-PCD (BM1407).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The study was approved by the Cyprus National Bioethics Committee (EEBK/EΠ/2013/21) and it conforms to the provisions of the Helsinki Declaration. All patients provided informed consent and patient anonymity was preserved.
References
- Knowles MR, Daniels LA, Davis SD, et al. Primary ciliary dyskinesia. Recent advances in diagnostics, genetics, and characterization of clinical disease. Am J Respir Crit Care Med 2013;188:913-22. [Crossref] [PubMed]
- Kuehni CE, Frischer T, Strippoli MP, et al. Factors influencing age at diagnosis of primary ciliary dyskinesia in European children. Eur Respir J 2010;36:1248-58. [Crossref] [PubMed]
- Lucas JS, Burgess A, Mitchison HM, et al. Diagnosis and management of primary ciliary dyskinesia. Arch Dis Child 2014;99:850-6. [Crossref] [PubMed]
- Collins SA, Gove K, Walker W, et al. Nasal nitric oxide screening for primary ciliary dyskinesia: systematic review and meta-analysis. Eur Respir J 2014;44:1589-99. [Crossref] [PubMed]
- Raidt J, Wallmeier J, Hjeij R, et al. Ciliary beat pattern and frequency in genetic variants of primary ciliary dyskinesia. Eur Respir J 2014;44:1579-88. [Crossref] [PubMed]
- Papon JF, Coste A, Roudot-Thoraval F, et al. A 20-year experience of electron microscopy in the diagnosis of primary ciliary dyskinesia. Eur Respir J 2010;35:1057-63. [Crossref] [PubMed]
- Pazour GJ, Agrin N, Leszyk J, et al. Proteomic analysis of a eukaryotic cilium. J Cell Biol 2005;170:103-13. [Crossref] [PubMed]
- Becker-Heck A, Loges NT, Omran H. Dynein dysfunction as a cause of primary ciliary dyskinesia and other ciliopathies. Dyneins: Elsevier, 2011:602-27.
- Davis SD, Ferkol TW, Rosenfeld M, et al. Clinical features of childhood primary ciliary dyskinesia by genotype and ultrastructural phenotype. Am J Respir Crit Care Med 2015;191:316-24. [Crossref] [PubMed]
- Marshall CR, Scherer SW, Zariwala MA, et al. Whole-Exome Sequencing and Targeted Copy Number Analysis in Primary Ciliary Dyskinesia. G3 (Bethesda) 2015;5:1775-81. [Crossref] [PubMed]
- Frommer A, Hjeij R, Loges NT, et al. Immunofluorescence analysis and diagnosis of primary ciliary dyskinesia with radial spoke defects. Am J Respir Cell Mol Biol 2015;53:563-73. [Crossref] [PubMed]
- Onoufriadis A, Shoemark A, Schmidts M, et al. Targeted NGS gene panel identifies mutations in RSPH1 causing primary ciliary dyskinesia and a common mechanism for ciliary central pair agenesis due to radial spoke defects. Hum Mol Genet 2014;23:3362-74. [Crossref] [PubMed]
- Alsaadi MM, Gaunt TR, Boustred CR, et al. From a single whole exome read to notions of clinical screening: primary ciliary dyskinesia and RSPH9 p. Lys268del in the Arabian Peninsula. Ann Hum Genet 2012;76:211-20. [Crossref] [PubMed]
- Castleman VH, Romio L, Chodhari R, et al. Mutations in radial spoke head protein genes RSPH9 and RSPH4A cause primary ciliary dyskinesia with central-microtubular-pair abnormalities. Am J Hum Genet 2009;84:197-209. [Crossref] [PubMed]
- Werner C, Onnebrink JG, Omran H. Diagnosis and management of primary ciliary dyskinesia. Cilia 2015;4:2. [Crossref] [PubMed]
- Sisson JH, Stoner J, Ammons B, et al. All‐digital image capture and whole‐field analysis of ciliary beat frequency. J Microsc 2003;211:103-11. [Crossref] [PubMed]
- Yiallouros PK, Kouis P, Middleton N, et al. Clinical features of primary ciliary dyskinesia in Cyprus with emphasis on lobectomized patients. Respir Med 2015;109:347-56. [Crossref] [PubMed]
- Van der Auwera GA, Carneiro MO, Hartl C, et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Curr Protoc Bioinformatics 2013;43:11.10.1-33.
- Quanjer PH, Stanojevic S, Cole TJ, et al. Multi-ethnic reference values for spirometry for the 3-95-yr age range: the global lung function 2012 equations. Eur Respir J 2012;40:1324-43. [Crossref] [PubMed]
- Kuczmarski RJ, Ogden CL, Guo SS, et al. 2000 CDC Growth Charts for the United States: methods and development. Vital Health Stat 11 2002;246:1-190. [PubMed]
- Ellard S, Baple EL, Owens M, et al. ACGS best practice guidelines for variants classification 2017 – consensus statement. Assoc Clin Genet Sci 2017;1-12.
- Fagerberg L, Hallström BM, Oksvold P, et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol Cell Proteomics 2014;13:397-406. [Crossref] [PubMed]
- Chilvers MA, Rutman A, O'Callaghan C. Ciliary beat pattern is associated with specific ultrastructural defects in primary ciliary dyskinesia. J Allergy Clin Immunol 2003;112:518-24. [Crossref] [PubMed]
- Leigh MW, Hazucha MJ, Chawla KK, et al. Standardizing nasal nitric oxide measurement as a test for primary ciliary dyskinesia. Ann Am Thorac Soc 2013;10:574-81. [Crossref] [PubMed]
- Sosnay PR, Raraigh KS, Gibson RL. Molecular Genetics of Cystic Fibrosis Transmembrane Conductance Regulator: Genotype and Phenotype. Pediatr Clin North Am 2016;63:585-98. [Crossref] [PubMed]
- Moskowitz SM, Chmiel JF, Sternen DL, et al. Clinical practice and genetic counseling for cystic fibrosis and CFTR-related disorders. Genet Med 2008;10:851. [Crossref] [PubMed]
- Kott E, Legendre M, Copin B, et al. Loss-of-function mutations in RSPH1 cause primary ciliary dyskinesia with central-complex and radial-spoke defects. Am J Hum Genet 2013;93:561-70. [Crossref] [PubMed]
- Knowles MR, Ostrowski LE, Leigh MW, et al. Mutations in RSPH1 cause primary ciliary dyskinesia with a unique clinical and ciliary phenotype. Am J Respir Crit Care Med 2014;189:707-17. [Crossref] [PubMed]
- Olbrich H, Cremers C, Loges NT, et al. Loss-of-function GAS8 mutations cause primary ciliary dyskinesia and disrupt the nexin-dynein regulatory complex. Am J Hum Genet 2015;97:546-54. [Crossref] [PubMed]
- Shapiro AJ, Leigh MW. Value of transmission electron microscopy for primary ciliary dyskinesia diagnosis in the era of molecular medicine: Genetic defects with normal and non-diagnostic ciliary ultrastructure. Ultrastruct Pathol 2017;41:373-85. [Crossref] [PubMed]
- Shoemark A, Moya E, Hirst RA, et al. High prevalence of CCDC103 p.His154Pro mutation causing primary ciliary dyskinesia disrupts protein oligomerisation and is associated with normal diagnostic investigations. Thorax 2018;73:157-66. [Crossref] [PubMed]
- Pifferi M, Bush A, Michelucci A, et al. Mannose-binding lectin 2 gene polymorphism and lung damage in primary ciliary dyskinesia. Pediatr Pulmonol 2015;50:179-86. [Crossref] [PubMed]
- Dizier MH, Nadif R, Margaritte-Jeannin P, et al. Interaction between the DNAH9 gene and early smoke exposure in bronchial hyperresponsiveness. Eur Respir J 2016;47:1072-81. [Crossref] [PubMed]
- Brennan ML, Schrijver I. Cystic fibrosis: a review of associated phenotypes, use of molecular diagnostic approaches, genetic characteristics, progress, and dilemmas. J Mol Diagn 2016;18:3-14. [Crossref] [PubMed]
- Zielenski J. Genotype and phenotype in cystic fibrosis. Respiration 2000;67:117-33. [Crossref] [PubMed]
- Werner C, Lablans M, Ataian M, et al. An international registry for primary ciliary dyskinesia. Eur Respir J 2016;47:849-59. [Crossref] [PubMed]
- Goutaki M, Maurer E, Halbeisen FS, et al. The international primary ciliary dyskinesia cohort (iPCD Cohort): methods and first results. Eur Respir J 2017;49:1601181. [Crossref] [PubMed]
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 1 cilia motility assessment. Asvide 2019;6:137. Available online: http://www.asvide.com/article/view/31923
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 2 cilia motility assessment. Asvide 2019;6:138. Available online: http://www.asvide.com/article/view/31924
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 3 cilia motility assessment. Asvide 2019;6:139. Available online: http://www.asvide.com/article/view/31925
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 4 cilia motility assessment. Asvide 2019;6:140. Available online: http://www.asvide.com/article/view/31926
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 5 cilia motility assessment. Asvide 2019;6:141. Available online: http://www.asvide.com/article/view/31927
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 6 cilia motility assessment. Asvide 2019;6:142. Available online: http://www.asvide.com/article/view/31928
- Yiallouros PK, Kouis P, Pirpa P, et al. Case 7 cilia motility assessment. Asvide 2019;6:143. Available online: http://www.asvide.com/article/view/31929


