Detection of plasma T790M mutation after the first generation EGFR-TKI resistance of non-small cell lung cancer in the real world
Introduction
With the increase of pollution and a growing smoking population, the incidence of lung cancer has increased (1). For most patients with advanced stage lung cancer, treatment is difficult, prognosis is poor, chemotherapy sensitivity is low, and the mortality rate is very high (2). Non-small cell lung cancer (NSCLC) accounts for about 80–85% of all pathological types. Along with the development of molecular detection technology, precision medicine requires individualized diagnosis and treatment. The epidermal growth factor receptor (EGFR) gene has been identified as the driving gene of non-small cell lung cancer. EGFR is the most common driver gene in East Asians, accounting for about 50% of cases. The EGFR gene mutation in patients with EGFR-TKI has a good curative effect, and the average progression-free survival (PFS) is 9–13 months (3,4). Although the majority EGFR-mutant NSCLC patients have a good response to EGFR-TKI initially, drug resistance is inevitable. It has been confirmed that there are many mechanisms of acquired resistance to EGFR-TKI, among which the secondary EGFR substitution of threonine to methionine at position 790 (T790M) mutation accounts for about 50% (5,6). The third-generation of EGFR-TKI has significant efficacy in advanced T790M-positive NSCLC patients (7,8). Therefore, it is necessary to clear the gene status of T790M in patients with acquired drug-resistance after the first generation of TKI drug treatment, as it can indicate the direction of subsequent treatment.
To detect if patients are harboring the T790M mutation, tissue biopsy specimens for molecular testing is accepted as the gold standard, but it is invasive and has several limitations, such as the occurrence of complications, poor quantity of sample and quality, repeat biopsy, and tumor heterogeneity. Liquid biopsy that collects plasmas pecimens offers a noninvasive approach that can be acquired in almost all clinical situations. Research shows it contains tumor genome information, and among them, cell-free DNA (cfDNA) is the most stable and easily available source (9). Liquid biopsy can be used for the diagnosis and evaluation of resistance mechanisms.
The objective of this study was to investigate the positive rate of plasma test T790M mutation and its relationship with different clinical characteristics and the frequency of T790M mutation in advanced EGFR-mutant NSCLC patients with acquired resistance after first line EGFR-TKI treatment.
Methods
Patients
This study was a prospective, single-center study at Taizhou Hospital in Zhejiang Province. We recruited patients from September 2017 to June 2018. The eligibility criteria included the following: (I) aged 18 years or older, with histologically confirmed NSCLC stage IIIB/st (10) with EGFR mutation positive in initial tissue detection, and who were unable to undergo operation or had recurrence after operation; (II) progressive disease (PD) after first generation EGFR-TKI by RECIST, with PFS >3 months confirmed by RESIST V1.1 (11); (III) no third generation TKI treatment.
EGFR mutation test
All patients signed the informed consent, and 10 mL of blood was taken to assess the presence of the T790M gene by amplification refractory mutation system (ARMS) according to the manufacturer’s instructions (Amoy Diagnostics Co., Ltd., China).
Statistical analyses
Univariate analysis by Chi-square test was performed for assessing the associations between patients’ characteristics, including age, sex, baseline EGFR mutation status, the type of first generation EGFR-TKI treatment, and the frequency of T790M mutation. The statistical tests were done with SPSS ver. 23.0 (IBM Corp., Armonk, NY, USA). P values of <0.05 were considered statistically significant. Plasma DNA was extracted by immunomagnetic beads.
Ethical statement
The study was approved by the Institutional Ethics Committee of Taizhou Hospital (ethical batch number: 201637).
Results
Patients’ characteristics
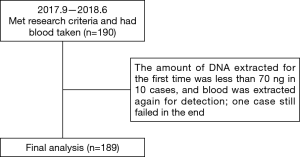
A total of 190 patients with locally advanced or advanced EGFR mutant lung adenocarcinoma met the research criteria, but one of them failed in the cfDNA collection. Thus, 189 patients were included in the analysis (Figure 1). The baseline characteristics are shown in Table 1. The average age of patients was 64.03 years old (range, 32–87 years old). There were 64 males (33.86%) and 125 females (66.14%). For baseline EGFR mutation status, 99 patients carried exon 19 deletion mutation (19del) (52.38%), 84 patients carried exon 21 L858R point mutation (L858R) (44.44%) and 6 patients carried mixed mutations (3.18%) (1 patient carried 19del + L858R, 1 patient carried L858R + S768I, 2 patients carried L858R+20-INS, 1 patient carried G719X + L861Q, and 1 patient carried L861Q+19del). For the first generation EGFR-TKI treatment type, 96 patients received icotinib (50.79%), 90 patients received gefitinib (47.62%), and 3 patients received erlotinib (1.59%). According to Wu et al.’s reports (12) of disease progression modes, 78 cases (41.27%) showed gradual progression, 48 cases (24.40%) showed local progression, and 63 cases (33.33%) showed dramatic progression.
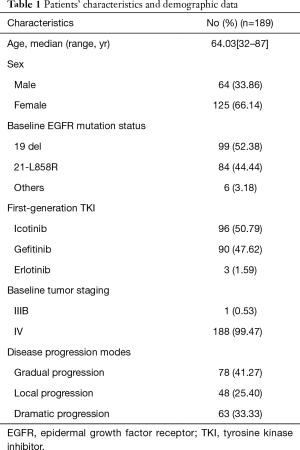
Full table
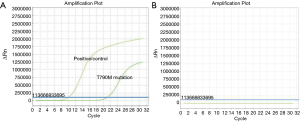
Positive rate of T790M mutation in plasma test
Judging from the results (Figure 2), of the 189 patients, 69 had T790M mutation detected in plasma, with a positive rate of 36.51%. The relationship between the mutation state and the detection of plasma T790M mutation is shown in Figure 3. According to the baseline EGFR gene mutation states, 45 of 99 patients with the 19del mutation had the T790M mutation (45.45%), 22 of 84 patients with L858R mutation had the T790M mutation (26.19%), and 2 of 6 patients with mixed mutations had the T790M mutation (33.33%).

The association between patients’ characteristics and the T790M mutation states of plasma detection
Univariate analyses of the association between patients’ characteristics and the T790M mutation states of blood plasma detection are summarized in Table 2. Among them, the patients with baseline 19del mutation had a significantly higher positive rate of T790M mutation than the ones who had L858R mutation and other mutations (P=0.026). For disease progression modes, the patients with local progression had a significantly higher positive rate of T790M mutation than the patients with gradual progression and dramatic progression (P=0.031). There were no relationships of T790M mutation status of plasma detection with age, sex, or the type of first generation EGFR-TKI treatment. Because only 1 patient was enrolled in stage IIIb, whether there was a relationship between disease staging and the T790M mutation state was not calculated.
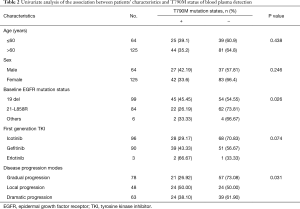
Full table
Discussion
With the development of molecular detection technology, it has been confirmed that the EGFR gene is the driving gene of lung cancer, and EGFR mutation positive is considered a predictor for EGFR-TKI efficacy. For NSCLC-EGFR mutation patients, EGFR-TKI has high efficiency and low toxicity side effects, but drug resistance is inevitable. Secondary EGFR mutation has been confirmed as the most common mechanism of acquired resistance to EGFR-TKI, with the T790M mutation accounting for about 50% of cases (4,6). In recent years, it has become clear that NSCLC patients with positive T790M mutation have a high response rate (about 60%) to third generation EGFR-TKI. For example, osimertinib, produced by AstraZeneca, can achieve an overall remission rate of 63% and a PFS of 9.7 months, which is considered effective treatment. Therefore, it is necessary to know the characteristics of NSCLC patients with T790M mutation after first generation EGFR-TKI treatment of acquired resistance. NCCN guidelines (13) suggest plasma detection to test the EGFR gene state after drug resistance, especially the T790M gene.
Tissue biopsies are invasive, and their possible association with complications and increased cost make them ill-suited for molecular testing. For patients with multiple metastases, different gene detection results may exist in different sampling sites (14). The above deficiencies can be well avoided by gene detection through plasma sampling. Genetic substances are released into circulation through several mechanisms, including apoptosis, necrosis, and exocytosis (15). Circulating tumor DNA (ctDNA) is part of cfDNA. Falling from tumor deposits into the vasculature, it is feasible to use peripheral blood for gene detection. Nakano et al. (16) found that the cfDNA in the plasma of lung cancer patients was higher than that of normal people. Meanwhile, Sundaresan et al. (17), when investigating the tumor tissue biopsy and non-invasive blood biopsy in patients, discovered that the genotyping result based on ctDNA was approximately equivalent to that based on tissue. Also, Douillard et al. (18) using synchronous detection of EGFR mutation by tumor tissue and plasma DNA, showed that the coincidence rate of plasma EGFR detection was 65.7%, and thus plasma testing can be used as a supplement to tissue specimens.
As it pertains to the detection method, Qin et al. (19) compared the methods of detecting EGFR mutations in plasma samples of patients with advanced NSCLC. In Scorpion Amplification Reference Mutation System (sARMS), denaturing high performance liquid chromatography (DHPLC), and direct sequencing method, the results showed that among 73 patients, the sARMS method detected 28 (38.4%) patients with EGFR mutations, direct DNA sequencing detected 5 of them (6.9%), and the DHPLC method detected 22 (30.1%). This shows that the sensitivity of sARMS is higher than DHPLC and direct sequencing method in detecting the EGFR gene mutation state of plasma DNA. Xu et al. (20) compared the methods of DHPLC, sARMS, and mutant-enriched PCR for testing EGFR mutations in peripheral plasma. The results showed that the mutation rates of EGFR in plasma were 15.7%, 29.4%, and 27.5%, respectively; thus, compared with the other two methods, sARM detection’s sensitivity was higher. The above data show that the sARMS method has a higher mutation sensitivity than DHPLC, the direct sequencing method, and the mutation enrichment PCR method.
In this study, the T790M mutation was detected by ARMS, and had a positive mutation rate of 36.51%, which is between the rate detected by ARMS as reported by Wei et al. (21) (30.6%) and that detected by ARMS as reported by Kuang et al. (22) (45.2%). The reason for this discrepancy may be due to different sensitivities of detection methods. Also, plasma DNA samples may contain mutations, wild-type ctDNA, and cfDNA of normal cell apoptosis. These factors will affect the detection positive rate of the plasma T790M mutation. The technology for how to acquire reliable DNA and continuously improve detection methods needs to be further explored in research.
The detection method of T790M mutation has continuously been updated and improved, both in terms of sensitivity and specificity. Next generation sequencing is currently the most advanced detection technology: not only is the sensitivity higher, but it can also provide more cancer gene information. However, it is both time-consuming and expensive, which limits its clinical application. In this study, we used ARMS to test for the T790M mutation, which had the advantages of simple and convenient operation, good repeatability, and widespread clinical use (23). Therefore, the ARMS method can be used to detect the T790M mutation state, and can partially replace the plasma detection of tissues.
Our study demonstrated that the frequency of T790M has no correlation with age, sex, and the type of first generation EGFR-TKI treatment. The EGFR 19del mutation at baseline and the mode of local progression after resistance to first generation EGFR-TKI were correlated with a higher frequency of the T790M mutation; thus, they can be a predictor of the T790M mutation.
Huang et al. (24) recruited 205 patients to explore the relationship of acquired T790M mutation with clinical characteristics after resistance to first generation EGFR-TKI in lung adenocarcinoma. They reported that the positive frequency of T790M mutation was higher in patients with EGFR 19del mutation than in those with EGFR L858R mutation and others mutations in baseline (55.0% vs. 37.3%vs. 27.3%, respectively; P=0.022; by Fisher’s Exact Test). Matsuo et al. (25) stated that the patients with EGFR 19del mutation in baseline had a higher frequency of the T790M mutation than patients with the L858R mutation (63% vs. 38% respectively; P=0.035). Nosaki et al. (26) also reported that the frequency of T790M mutation in patients with EGFR 19del mutation was higher than that in patients with L858R mutation (55.6% vs. 43.0% respectively; P=0.05). In addition, Huang et al. (24) demonstrated that patients with Del E746-A750 mutation of the subtypes of EGFR 19del had higher T790M mutation frequency than patients with other subtypes. In this study, the frequency of T790M mutation in patients with EGFR 19del mutation in baseline was significantly higher than that in patients with L858R and other mutations (P=0.026), which is similar to conclusions in other research reports. Six patients initially had complicated EGFR mutations in this study, and the frequency of T790M mutation was also lower than that of patients with initial 19del mutation. However, the sample size was small, and we need to expand it in further study.
Oya et al. (27) explored the relationship between the T790M mutation state and progression modes in EGFR TKI therapy. They found that the frequency of T790M mutation was higher in the patients with solitary lesion progression than in patients with multiple lesion progression (58% vs. 24% respectively; P<0.0001). Wei et al. (21) reported that the T790M mutation was related to disease progression mode, and that the local progression patients were more likely harbor the T790M mutation than those with distant metastasis (P=0.009). Our study also demonstrated that the T790M mutation was related to the progression mode of disease, and the frequency of T790M was higher in patients with local progression after EGFR-TKI acquired resistance than that in patients of gradual progression and dramatic progression (P=0.031). There is other research (28) that indicates the disease progression modes after first generation EGFR-TKI treatment are related to clinical results: solitary progression had significantly longer PFS than multi-site progression. This indicates that the growth of tumor cells progressing at solitary sites is more inert.
Cells sensitive to EGFR-TKI due to 19del mutation and acquired resistance due to T790M mutation proliferate in vitro at a slower rate than parent cells (29). The growth speed of acquired drug-resistant tumor cells caused by T790M mutation in vivo is also significantly slower than that of their parent cells. This partly explains why the T790M mutation frequency of patients with local progression is higher than that of other progression modes. As our study did not carry out tissue biopsy for control study, false-negatives could not be avoided in the plasma detection of T790M mutation; at the same time, due to the limitation of sample size, it is difficult to clarify the exact mechanism whether the mutation frequency of T790M in patients with local progression was higher than that of other progression modes, and further research is needed.
Our study had some limitations. First, this is a single-center study, and there might have been selection bias; second, the sample size of this study was not large enough, and no control tissue biopsy was performed. This study found that patients with 19 del deletion and local progression after TKI treatment may be the predictive factors of T790M mutation, but the molecular mechanism behind this still needs further exploration and research.
Acknowledgments
Funding: None.
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was approved by the Institutional Ethics Committee of Taizhou Hospital (ethical batch number: 201637).
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Minnix JA, Karam-Hage M, Blalock JA, et al. The importance of incorporating smoking cessation into lung cancer screening. Transl Lung Cancer Res 2018;7:272-80. [Crossref] [PubMed]
- Pereira JL, Gomes M, Teixeira AL, et al. Potential and importance of metalloproteinases and interleukins in inflammation and metastasization in non-small cell lung cancer. Transl Cancer Res 2018;7:782-95. [Crossref]
- Shepherd FA, Rodrigues Pereira J, Ciuleanu T, et al. Erlotinib in previously treated non-small-cell lung cancer. N Engl J Med 2005;353:123-32. [Crossref] [PubMed]
- Yoshida K, Kanda S, Shiraishi H, et al. Difference in central nerve system metastasis during gefitinib or erlotinib therapy in patients with EGFR-mutated non-small cell lung cancer: a retrospective study. J Thorac Dis 2019;11:1347-54. [Crossref] [PubMed]
- Yu HA, Arcila ME, Rekhtman N, et al. Analysis of tumor specimens at the time of acquired resistance to EGFR-TKI therapy in 155 patients with EGFR-mutant lung cancers. Clin Cancer Res 2013;19:2240-7. [Crossref] [PubMed]
- Karachaliou N, Fernandez-Bruno M, Bracht JW, et al. EGFR first- and second-generation TKIs—there is still place for them in EGFR-mutant NSCLC patients. Transl Cancer Res 2019;8:S23-47. [Crossref]
- Papadimitrakopoulou V, Wu YL, Ahn MJ, et al. PL03.03: Randomized Phase III Study of Osimertinib vs Platinum-Pemetrexed for EGFR T790M-Positive Advanced NSCLC (AURA3). J Thorac Oncol 2017;12:S5-6. [Crossref]
- Ma L, Chen R, Wang F, et al. EGFR L718Q mutation occurs without T790M mutation in a lung adenocarcinoma patient with acquired resistance to osimertinib. Ann Transl Med 2019;7:207. [Crossref] [PubMed]
- Forshew T, Murtaza M, Parkinson C, et al. Noninvasive identification and monitoring of cancer mutations by targeted deep sequencing of plasma DNA. Sci Transl Med 2012;4:136ra68. [Crossref] [PubMed]
- Edge SB, Compton CC. The American Joint Committee on Cancer: the 7th edition of the AJCC cancer staging manual and the future of TNM. Ann Surg Oncol 2010;7:1471-4.
- Eisenhauer EA, Therasse P, Bogaerts J, et al. New response evaluation criteria in solid tumours: revised RECIST guideline (version 1.1). Eur J Cancer 2009;45:228-47. [Crossref] [PubMed]
- Yang JJ, Chen HJ, Wu YL, et al. Clinical modes of EGFR tyrosine kinase inhibitor failure and subsequent management in advanced non-small cell lung cancer. Lung Cancer 2013;79:33-9. [Crossref] [PubMed]
- Ettinger DS, Wood DE, Aisner DL, et al. Non-Small Cell Lung Cancer, Version 5.2017, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw 2017;15:504-35. [Crossref] [PubMed]
- Piotrowska Z, Niederst MJ, Mino-Kenudson M, et al. Variation in mechanisms of acquired resistance among EGFR-mutant NSCLC patients with more than 1 post-resistant biopsy. Int J Radiat Oncol 2014;90:S6-7. [Crossref]
- Schwarzenbach H, Hoon DS, Pantel K. Cell-free nucleic acids as biomarkers in cancer patients. Nat Rev Cancer 2011;11:426-37. [Crossref] [PubMed]
- Nakano H, Soda H, Takasu M, et al. Heterogeneity of epidermal growth factor receptor mutations within a mixed adenocarcinoma lung nodule. Lung Cancer 2008;60:136-40. [Crossref] [PubMed]
- Sundaresan TK, Sequist LV, Heymach JV, et al. Detection of T790M, the acquired resistance EGFR mutation, by tumor biopsy versus noninvasive blood-based analyses. Clin Cancer Res 2016;22:1103-10. [Crossref] [PubMed]
- Douillard JY, Ostoros G, Cobo M, et al. First-line gefitinib in Caucasian EGFR mutation-positive NSCLC patients: a phase-IV, open-label, single-arm study. Br J Cancer 2014;110:55-62. [Crossref] [PubMed]
- Qin L, Zhong W, Zhang L, et al. Comparison of three methods for detecting epidermal growth factor receptor mutations in plasma DNA samples of Chinese patients with advanced non-small cell lung cancer. Chin Med J 2011;124:887-91. [PubMed]
- Xu F, Wu J, Xue C, et al. Comparison of different methods for detecting epidermal growth factor receptor mutations in peripheral blood and tumor tissue of non-small cell lung cancer as a predictor of response to gefitinib. Onco Targets Ther 2012;5:439-47. [PubMed]
- Wei Y, Wei L, Ma XP, et al. Correlation with advanced NSCLC T790M EGFR-TKI secondary resistance and prognosis. Chin J Cancer Prev Treat 2016;23:364-8.
- Kuang Y, Rogers A, Yeap BY, et al. Noninvasive detection of EGFR T790M in gefitinib or erlotinib resistant non-small cell lung cancer. Clin Cancer Res 2009;15:2630-6. [Crossref] [PubMed]
- Zhang H, Yang XJ, Zhang XY, et al. Detection and analysis of EGFR mutation with advanced lung adenocarcinoma. Chin J Cancer PrevTreat 2014;21:1517-20.
- Huang YH, Hsu KH, Tseng JS, et al. The Association of Acquired T790M Mutation with Clinical Characteristics after Resistance to First-Line Epidermal Growth Factor Receptor Tyrosine Kinase Inhibitor in Lung Adenocarcinoma. Cancer Res Treat 2018;50:1294-303. [Crossref] [PubMed]
- Matsuo N, Azuma K, Sakai K, et al. Association of EGFR exon 19 deletion and EGFR-TKI treatment duration with frequency of T790M mutation in EGFR-mutant lung cancer patients. Sci Rep 2016;6:36458. [Crossref] [PubMed]
- Nosaki K, Satouchi M, Kurata T, et al. Re-biopsy status among non-small cell lung cancer patients in Japan: a retrospective study. Lung Cancer 2016;101:1-8. [Crossref] [PubMed]
- Oya Y, Yoshida T, Kuroda H, et al. Association Between EGFR T790M Status and Progression Patterns During Initial EGFR-TKI Treatment in Patients Harboring EGFR Mutation. Clin Lung Cancer 2017;18:698-705.e2. [Crossref] [PubMed]
- Yoshida T, Yoh K, Niho S, et al. RECIST progression patterns during EGFR tyrosine kinase inhibitor treatment of advanced non-small cell lung cancer patients harboring an EGFR mutation. Lung Cancer 2015;90:477-83. [Crossref] [PubMed]
- Chmielecki J, Foo J, Oxnard GR, et al. Optimization of dosing for EGFR-mutant non-small cell lung cancer with evolutionary cancer modeling. Sci Transl Med 2011;3:90ra59. [Crossref] [PubMed]