Serum exosomal microRNA-146a as a novel diagnostic biomarker for acute coronary syndrome
Introduction
Acute coronary syndrome (ACS), which includes ST-segment elevation myocardial infarction (STEMI), non-ST-segment elevation myocardial infarction (NSTEMI), and unstable angina (UA), is a clinical syndrome of myocardial ischemia caused by rupture of unstable coronary plaque and thrombosis (1,2) and is characterized by acute onset, fast progression, and extremely high mortality rate. Early diagnosis and treatment of ACS is of great significance to improve therapeutic effects. Studies have found that inflammation plays a vital role in the pathogenesis of ACS (3). The inflammatory response activated by an ACS attack can lead to the imbalance of adipokine and can be ameliorated by anti-inflammation drugs (4). Westman et al. (5) confirmed that inflammatory response is the driving factor of adverse left ventricular remodeling after ACS. Therefore, to exploration of the relevant mechanisms of the regulation of inflammatory response in ACS is necessary.
Studies have reported that cells can actively secrete exosomes through exocytosis to mediate cell-to-cell communication. Exosomes can transfer a large number of proteins, RNA, microRNA (miRNA), and other substances from donor cells to recipient cells, thereby participating in the transmission of intercellular information (6). Almost all cells can secrete exosomes, including stem cells, endothelial cells, epithelial cells, dendritic cells, and so on (7-9). Exosomes have been widely found in various body fluids, such as blood, urine, amniotic fluid, and pleural fluid. In human blood, the concentration of exosomes is about 1010/mL (10). Exosomes protect the miRNA existing in them from degradation and ensure the stable circulation of miRNA in the blood, so that miRNA can transfer between cells and exert regulatory effects. The expression levels of miR-208a in serum exosomes of ACS patients was noticeably higher than in healthy controls. Kaplan-Meier survival analysis results have shown that 1-year survival rate of patients with high expression levels of serum exo-miR-208a was significantly reduced, indicating that a high level of serum exo-miR-208a was associated with poor prognosis of ACS (11). Ling et al. (12) found that serum exo-miR-21 and exo-miR-126 might be used as biomarkers for ACS diagnosis.
The miRNA miR-146a, which is located on human chromosome 5q33, has a protective effect on apoptosis induced by myocardial ischemia/hypoxia (13), and is also highly expressed in ACS patients (14,15). It has been shown through receiver operating characteristic (ROC) curve analysis that miR-146a has a high diagnostic efficiency in predicting acute myocardial infarction (MI) (16), and the silent miR-146a can also alleviate heart failure in rats with MI (17).
We conducted this study to explore whether serum exo-miR-146a could be used as a novel diagnostic biomarker for ACS and to investigate its relationship with inflammatory response. We present the following article in accordance with the STARD reporting checklist (available at http://dx.doi.org/10.21037/jtd-21-609).
Methods
Clinical samples
Patients aged 40–80 years old who visited the First Affiliated Hospital of Kunming Medical University for treatments due to “chest pain” from May 2019 to December 2020 were enrolled prospectively. A total of 21 patients with STEMI, 23 patients with NSTEMI, and 19 patients with UA were consecutively included in this study. At the same time, 25 patients with normal coronary arteries confirmed by coronary angiography were selected as the control group. The medical history, laboratory examination, and echocardiographic results were collected after admission. Each study was conducted after the participants voluntarily signed an informed consent form.
The diagnosis of ACS was based on the guideline of the European Society of Cardiology. The exclusion criteria were as follows: (I) patients with malignant tumors, immune system diseases, cerebral vascular diseases, peripheral vascular disease, and blood system diseases; (II) patients with recent high fever, complicated with acute or chronic viral and bacterial infection; (III) heart valve disease, cardiomyopathy, congenital heart disease, cardiac shock; (IV) a history of MI, patients who had undergone previous cardiac surgical procedures.
The study protocol was approved by the Ethics Committee of the First Affiliated Hospital of Kunming Medical University (No. 2020-L-17), which was conducted according to the Declaration of Helsinki (as revised in 2013).
Blood samples
For participants undergoing primary percutaneous coronary intervention (pPCI), 5 mL venous blood was collected by dipotassium ethylenediaminetetraacetic acid (EDTA-K2) anticoagulant tube immediately after admission. For participants without emergency surgery, the blood sample was collected at 7:00 am before breakfast. After centrifugation at 4 °C and 3,000 g for 15 min, the supernatant was collected into an RNA-free EP tube and stored in a refrigerator at −80 °C for later use.
Serum exosomes isolation
The collected serum was centrifuged at 3,000 g and 4 °C for 15 minutes to remove cells and cell debris. Then, 250 µL of supernatant was transferred to a sterile container, and 63 µL Exoquick precipitation solution (Systems Biosciences, Palo Alto, CA, USA) was added. After refrigeration at 4 °C for 30 min, the mixture was centrifuged at 1,500 g and 4 °C for 30 min. The supernatant (exosome-free) and white precipitate (exosome) were collected, respectively. The precipitation was resuspended with 250 µL phosphate-buffered saline (PBS).
Transmission electron microscope
A total of 10 µL exosome suspension solution was dropped on the special copper grid and allowed to stand for 1 min at room temperature. The excess liquid was absorbed from the edge of the copper grid with a clean filter paper. A 10 g/L phosphotungstate solution with pH 6.8 was re-stained and dried at room temperature for 5 min. Imaging observation and photography were conducted at 80–120 kV by cryo transmission electron microscopy (TEM, HT-7700; Hitachi, Tokyo, Japan), and the exospheric diameters were calculated with scales.
Particle size analysis
Particle size analysis of exosomes was performed by nanoparticle tracking analysis (NTA, NanoFCM, Beijing, China). The exosomes were diluted 50-fold in PBS; light scattering observation was performed with a conventional optical microscope perpendicular to the beam axis. The NTA software was used to track the Brownian motion of a single vesicle between frames and the Stokes Einstein equation was used to calculate the total concentration and magnitude.
Western blotting
Precipitation of the exosomes was resuspended in radioimmunoprecipitation assay (RIPA) buffer (Solarbio, Beijing, China) to extract exosome marker proteins, and then the protein concentration was determined by bicinchoninic acid (BCA, Thermo Fisher Scientific, Waltham, MA, USA). After sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE), the protein was transferred to a polyvinylidene fluoride (PVDF) membrane, sealed by 5% skimmed milk powder at room temperature, and then the exosome specific primary antibody against CD9, CD63, CD81, and HSP70 (1:1,000, System Biosciences) was incubated at 4 °C overnight. The primary antibody was discarded next day, then goat anti-Rabbit horseradish peroxidase (HRP, 1:20,000, System Biosciences) was added and incubated at 37 °C for 1 h. After incubating the blot with chemiluminescent substrate, it was exposed and photographed on a gel imager.
Enzyme-linked immunosorbent assay (ELISA)
The expression levels of interleukin-1β (IL-1β), interleukin-6 (IL-6), and tumor necrosis factor-α (TNF-α) in serum were determined by commercially available ELISA kits (4A Biotech, Beijing, China). After the cytokine antibody was diluted, it was added to the plate and then incubated at 37 °C for 2 h. The solution was sealed at 37 °C for 1 h, then the sample was added to the plate and incubated overnight at 4 °C. After washing with PBS, HRP-labeled secondary antibodies were added and incubated at 37 °C for 1 h the next day. The 3,3',5,5'-Tetramethylbenzidine (TMB) substrate chromo-developing solution was added, and after 15 min of color development at 37 °C, 2 mol/L sulfuric acid was added, the optical density (OD)450 value was detected on the microplate reader, and the blank control well was zeroed.
Real-time quantitative polymerase chain reaction (RT-qPCR)
A TRIzol LS reagent (Thermo Fisher Scientific) was used to extract total RNA from serum in each group according to the manufacturer’s protocol. After the concentration was determined, All-in-One miRNA qRT-PCR Detection Kit (GeneCopoeia, Rockville, MD, USA) was used to reverse transcribe the extracted total RNA into cDNA, and quantitative polymerase chain reaction (qPCR) was performed following the manufacturer’s protocol. The reaction procedure was as follows: pre-denaturation at 95 °C for 5 min, denaturation at 95 °C for 5 s, annealing at 60 °C for 30 s, elongation at 72 °C for 15 s, and 40 cycles. With U6 as the internal reference, the 2−∆∆Ct method was used to quantify miR-146a.
Statistical analysis
The data was analyzed using the software GraphPad Prism 7.0 (GraphPad software, San Diego, CA, USA) and SPSS version 20 (IBM, Armonk, NY, USA). One-way analysis of variance (ANOVA, parametric data) and non-parametric test (non-parametric data) were used for comparison between ACS patients and the controls. Spearman’s correlation analysis was performed to assess the correlation between serum exo-miR-146a and potential factors. The ROC curve analyses were performed to assess the predictive accuracy via area under curve (AUC). A P value <0.05 indicated a statistically significant difference.
Results
Comparison of clinical data
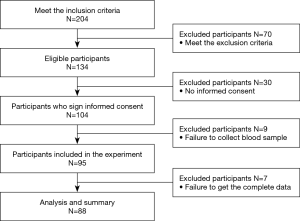
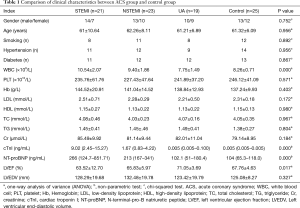
Figure 1 shows the flow diagram of participants selected. A total of 25 participants with normal coronary arteries confirmed by coronary angiography, 21 participants with STEMI, 23 participants with NSTEMI, and 19 participants with UA were enrolled in this study. According to Table 1, there were statistically significant differences in white blood cell (WBC), cardiac troponin (cTnI), N-terminal pro-B-type natriuretic peptide (NT-proBNP), and left ventricular ejection fraction (LVEF) among the 4 groups, while no other differences were found.
Full table
Characterization of exosomes
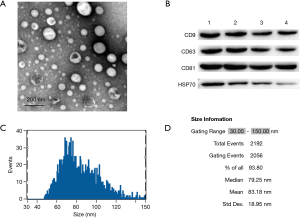
The morphological features of exosomes were observed by TEM, as shown in Figure 2A. The exosomes were of a typical cup-like shape. Meanwhile, exosomes specific markers CD9, CD63, CD81, and HSP70 were found to be positive by western blot (WB) in 4 random samples (Figure 2B). The results of particle size analysis showed that the exosomes with diameters of 30–150 nm accounted for 93.80%. In addition, the average diameter of exosomes was 83.18 nm, the median diameter was 79.25 nm, and the standard error (SE) was 18.95 nm (Figure 2C). The above results indicated that the serum exosomes isolated from serum could be used for the study of subsequent experiments.
The expression levels of exo-miR-146a in different groups
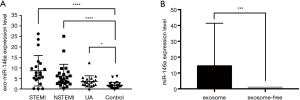
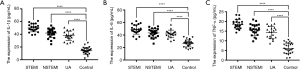
The expression level of miR-146a in exosomes was detected by RT-qPCR, and the results showed that miR-146a was dramatically increased in serum exosome of ACS patients compared with the control group (STEMI vs. control, P<0.001; NSTEMI vs. control, P<0.001; UA vs. control, P<0.05, Figure 3A). Interestingly, the expression of miR-146a in exosomes was significantly higher than exosome-free supernatant (P<0.05, Figure 3B).
The expression levels of IL-1β, IL-6, and TNF-α in different groups
We used ELISA to assess the expression of inflammation-related factors in the control group and ACS participants. As shown in Figure 4A,B,C, the expression of IL-1β, IL-6, and TNF-α in STEMI participants, NSTEMI participants, and UA participants were also significantly higher than the control group (P<0.0001, both).

Correlation analyses
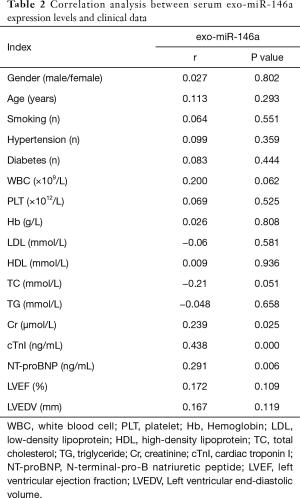
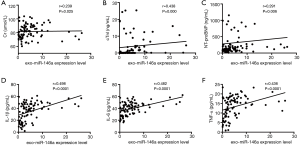
Spearman’s correlation analysis was performed to assess the potential factors related to the expression levels of serum exo-miR-146a. According to the results displayed in Table 2 and Figure 5A,B,C,D,E,F, exo-miR-146a expression was markedly correlated with creatinine (Cr, r=0.239, P=0.025), cTnI (r=0.438, P=0.000), NT-proBNP (r=0.291, P=0.006), IL-1β (r=0.498, P<0.0001), IL-6 (r=0.482, P<0.0001), and TNF-α (r=0.436, P<0.0001).
Full table
ROC curve analyses
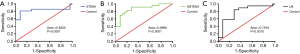
We performed ROC curve analysis to assess the predictive accuracy for ACS participants via AUC. As shown in Figure 6A,B,C, the AUC of serum exo-miR-146a was 0.8305 [SE =0.0741; 95% confidence interval (CI): 0.6853 to 0.9756; P=0.0001] in STEMI group, 0.8696 (SE =0.0529; CI: 0.7659 to 0.9732; P<0.0001) in NSTEMI group, and 0.7916 (SE =0.0719; CI: 0.6506 to 0.9326; P=0.0010) in UA group.

Discussion
The term ACS refers to the acute ischemic syndrome of the heart caused by the rupture or erosion of unstable atherosclerotic plaque in the coronary arteries. Studies have shown that inflammatory response is involved in the whole process of the occurrence and development of ACS. Myocardial ischemia and reperfusion can promote the production of IL-1β, IL-6, TNF-α, and so on (18,19). Inflammatory response is closely related to the development and prognosis of ACS (20). In recent years, it has been found that exosomes were widely participating in the physiological and pathological processes of the cardiovascular system, and play an important role in the evolution of heart diseases (21). Furthermore, it was found that exosomes carried out intercellular communication via secreting proteins, miRNAs, lncRNAs, and other contents (22). For example, the exosome miR-93-5p derived from adipose-derived stromal cells has a protective effect on myocardial injury induced by acute MI (23). Zhu et al. (24) demonstrated that bone marrow mesenchymal stem cells can prevent myocardial cell death during MI and promote cardiac recovery by secreting the exosome miR-125. Moreover, exosomes were found widely in various body fluids, such as blood, urine, amniotic fluid, and pleural fluid (25). Serum exosome miR-143 was under-expressed in patients with myocardial ischemia-reperfusion injury, and the overexpression of miR-143 inhibited angiogenesis (26). The expression level of miR-208 in serum exosomes of ACS patients was significantly higher than that of healthy controls and was associated with the prognosis of ACS (11).
In this study, exosomes of disk-like vesicles with a diameter of 30–150 nm were successfully isolated from the serum of ACS participants and the control group. The results of RT-qPCR showed that miR-146a was highly expressed in serum exosomes of ACS patients, and inflammatory factors (IL-1β, IL-6, and TNF-α) were also increased. Furthermore, Spearman’s correlation analysis results confirmed that the expression of serum exo-miR-146a was related to IL-1β, IL-6, and TNF-α, indicating that the high expression level of serum exo-miR-146a might be related to the inflammatory response.
Interestingly, the study also found that the serum level of miR-146a was mainly expressed in exosomes, which is consistent with Shimasaki’s study, which found that exosomes can ensure the stable circulation of miRNA in the blood.
Analysis of the ROC curves substantiated that exo-miR-146a might be a diagnostic biomarker for STEMI, NSTEMI, and UA, with good predictive accuracy to distinguish ACS patients from healthy people. Xue et al. (16) reported that plasma miR-146a was highly expressed in patients with MI, and ROC curves analysis showed that miR-146a had a higher diagnostic efficiency in predicting MI. He et al. (17) found that silencing miR-146a could alleviate heart failure in rats with MI. Our results are consistent with those of other scholars.
However, some limitations of this study need to be considered. First, the sample size of this study was small and the follow-up data was relatively absent, further studies are required with a larger sample size and longer follow-up time. Second, the specific mechanism of exo-miR-146a and inflammatory response is still unclear, the depth of research should be increased through cell experiments and animal model studies.
In conclusion, the serum exo-miR-146a may be used as a novel diagnostic biomarker for ACS patients, and it is also associated with inflammatory response.
Acknowledgments
Funding: This work was supported by the National Natural Science Foundation of China (81860073, 81860074, 81560075); the Applied and Basic Research Foundation of Yunnan Provincial Science and Technology Commission (2018FE001-(005); Foundation Projects of Yunnan Provincial Department of Education (2018JS206).
Footnote
Reporting Checklist: The authors have completed the STARD reporting checklist. Available at http://dx.doi.org/10.21037/jtd-21-609
Data Sharing Statement: Available at http://dx.doi.org/10.21037/jtd-21-609
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jtd-21-609). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study protocol was approved by the Ethics Committee of the First Affiliated Hospital of Kunming Medical University (No. 2020-L-17), which was conducted according to the Declaration of Helsinki (as revised in 2013). Each study was conducted after the participants voluntarily signed an informed consent form.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
Reference
- Switaj TL, Christensen SR, Brewer DM. Acute Coronary Syndrome: Current Treatment. Am Fam Physician 2017;95:232-40. [PubMed]
- Gach O, El HZ, Lancellotti P. Acute coronary syndrome. Rev Med Liege 2018;73:243-50. [PubMed]
- Libby P, Tabas I, Fredman G, et al. Inflammation and its resolution as determinants of acute coronary syndromes. Circ Res 2014;114:1867-79. [Crossref] [PubMed]
- Li R, Chen LZ, Zhao SP, et al. Inflammation Activation Contributes to Adipokine Imbalance in Patients with Acute Coronary Syndrome. PLoS One 2016;11:e0151916 [Crossref] [PubMed]
- Westman PC, Lipinski MJ, Luger D, et al. Inflammation as a Driver of Adverse Left Ventricular Remodeling After Acute Myocardial Infarction. J Am Coll Cardiol 2016;67:2050-60. [Crossref] [PubMed]
- Shimasaki T, Yamamoto S, Arisawa T. Exosome Research and Co-culture Study. Biol Pharm Bull 2018;41:1311-21. [Crossref] [PubMed]
- Gong M, Yu B, Wang J, et al. Mesenchymal stem cells release exosomes that transfer miRNAs to endothelial cells and promote angiogenesis. Oncotarget 2017;8:45200-12. [Crossref] [PubMed]
- Gao W, Liu H, Yuan J, et al. Exosomes derived from mature dendritic cells increase endothelial inflammation and atherosclerosis via membrane TNF-α mediated NF-κB pathway. J Cell Mol Med 2016;20:2318-27. [Crossref] [PubMed]
- Gupta R, Radicioni G, Abdelwahab S, et al. Intercellular Communication between Airway Epithelial Cells Is Mediated by Exosome-Like Vesicles. Am J Respir Cell Mol Biol 2019;60:209-20. [Crossref] [PubMed]
- Vicencio JM, Yellon DM, Sivaraman V, et al. Plasma exosomes protect the myocardium from ischemia-reperfusion injury. J Am Coll Cardiol 2015;65:1525-36. [Crossref] [PubMed]
- Bi S, Wang C, Jin Y, et al. Correlation between serum exosome derived miR-208a and acute coronary syndrome. Int J Clin Exp Med 2015;8:4275-80. [PubMed]
- Ling H, Guo Z, Shi Y, et al. Serum Exosomal MicroRNA-21, MicroRNA-126, and PTEN Are Novel Biomarkers for Diagnosis of Acute Coronary Syndrome. Front Physiol 2020;11:654. [Crossref] [PubMed]
- Huang W, Tian SS, Hang PZ, et al. Combination of microRNA-21 and microRNA-146a Attenuates Cardiac Dysfunction and Apoptosis During Acute Myocardial Infarction in Mice. Mol Ther Nucleic Acids 2016;5:e296 [Crossref] [PubMed]
- Bao MH, Xiao Y, Zhang QS, et al. Meta-Analysis of miR-146a Polymorphisms Association with Coronary Artery Diseases and Ischemic Stroke. Int J Mol Sci 2015;16:14305-17. [Crossref] [PubMed]
- Roldán V, Arroyo AB, Salloum-Asfar S, et al. Prognostic role of MIR146A polymorphisms for cardiovascular events in atrial fibrillation. Thromb Haemost 2014;112:781-8. [Crossref] [PubMed]
- Xue S, Zhu W, Liu D, et al. Circulating miR-26a-1, miR-146a and miR-199a-1 are potential candidate biomarkers for acute myocardial infarction. Mol Med 2019;25:18. [Crossref] [PubMed]
- He J, Lu Y, Song X, et al. Inhibition of microRNA-146a attenuated heart failure in myocardial infarction rats. Biosci Rep 2019;39:BSR20191732 [Crossref] [PubMed]
- Caruso R, Rocchiccioli S, Gori AM, et al. Inflammatory and antioxidant pattern unbalance in "clopidogrel-resistant" patients during acute coronary syndrome. Mediators Inflamm 2015;2015:710123 [Crossref] [PubMed]
- Anzai T. Inflammatory Mechanisms of Cardiovascular Remodeling. Circ J 2018;82:629-35. [Crossref] [PubMed]
- Huang WC, Chou RH, Chang CC, et al. Systemic Inflammatory Response Syndrome is an Independent Predictor of One-Year Mortality in Patients with Acute Myocardial Infarction. Acta Cardiol Sin 2017;33:477-85. [PubMed]
- Sahoo S, Losordo DW. Exosomes and cardiac repair after myocardial infarction. Circ Res 2014;114:333-44. [Crossref] [PubMed]
- Mathivanan S, Fahner CJ, Reid GE, et al. ExoCarta 2012: database of exosomal proteins, RNA and lipids. Nucleic Acids Res 2012;40:D1241-4. [Crossref] [PubMed]
- Liu J, Jiang M, Deng S, et al. miR-93-5p-Containing Exosomes Treatment Attenuates Acute Myocardial Infarction-Induced Myocardial Damage. Mol Ther Nucleic Acids 2018;11:103-15. [Crossref] [PubMed]
- Zhu LP, Tian T, Wang JY, et al. Hypoxia-elicited mesenchymal stem cell-derived exosomes facilitates cardiac repair through miR-125b-mediated prevention of cell death in myocardial infarction. Theranostics 2018;8:6163-77. [Crossref] [PubMed]
- Ibrahim A, Marbán E. Exosomes: Fundamental Biology and Roles in Cardiovascular Physiology. Annu Rev Physiol 2016;78:67-83. [Crossref] [PubMed]
- Geng T, Song ZY, Xing JX, et al. Exosome Derived from Coronary Serum of Patients with Myocardial Infarction Promotes Angiogenesis Through the miRNA-143/IGF-IR Pathway. Int J Nanomedicine 2020;15:2647-58. [Crossref] [PubMed]
(English Language Editor: J. Jones)



