
LncRNA SNHG7 promotes non-small cell lung cancer progression and cisplatin resistance by inducing autophagic activity
Highlight box
Key findings
• SNHG7 may have value as a novel biomarker of poor DDP responsiveness in NSCLC patients and may serve as a promising target for therapeutic intervention in these patients.
What is known and what is new?
• SNHG7 has recently been reported to be upregulated in breast, gastric, prostate, and colorectal cancers. Importantly, SNHG7 has been shown to promote NSCLC development.
• The role of SNHG7 as a regulator of autophagy in DDP-resistant NSCLC has not yet been established. Thus, the present study was developed to explore the role of SNHG7 in DDP-resistant NSCLC and to clarify the underlying mechanisms regulating this activity.
What are the implications, and what should change now?
• SNHG7 can promote malignant behaviors and DDP resistance in NSCLC cells in part via the induction of autophagic activity. However, the specific underlying mechanisms linking the regulation of SNHG7 to autophagic activity warrant further investigation.
Introduction
Lung cancer is one of the most prevalent and deadliest cancers globally (1). Broadly, lung cancer cases are separated into relatively rare small-cell lung cancers and the more common non-small cell lung cancers (NSCLCs), which account for 85–90% of the total caseload (2,3). Although many advances in the treatment and diagnosis of this condition have been made in recent years, NSCLC patients still face relatively poor 5-year overall survival (OS) outcomes (4,5). Platinum-based dual chemotherapy remains the standard approach to treating patients with advanced-stage disease (6,7). The emergence of chemoresistance, however, remains a major barrier to successful patient treatment (8). More comprehensive approaches to exploring the underlying basis for such chemoresistance are thus required to effectively treat affected patients.
Long non-coding RNAs (lncRNAs) are important regulators of a range of oncogenic processes including proliferative activity, differentiation, and development (9,10). For example, Luo et al. found that the lncRNA TRERNA1 targets FOXL1, thereby promoting NSCLC malignancy (11). Other groups have observed that lncRNAs act as mediators of chemoresistance, as in the case of CASC9, which is a lncRNA that has been found to promote epigenetic DUSP1 repression, thereby promoting NSCLC cell resistance to gefitinib treatment (12). Moreover, the lncRNA SNHG1 can target the miR-330-5p/DCLK1 axis to support cisplatin (DDP) resistance in NSCLC cells (13). However, the full spectrum of lncRNAs capable of influencing chemosensitivity-related processes has yet to be fully established.
SNHG7 is a recently identified 2,176 bp lncRNA encoded on chromosome 9q34.3 (14) and is reportedly upregulated in breast, gastric, prostate, and colorectal tumors (15). A growing body of evidence supports the ability of SNHG7 to drive tumor cell proliferative, migratory, and invasive activity, while at the same time protecting these malignant cells against apoptotic death (16-18). The overexpression of SNHG7 is also correlated with poorer clinicopathological characteristics and worse patient outcomes, suggesting that it may offer value as a prognostic and/or diagnostic biomarker, in addition to serving as a viable target for therapeutic anticancer treatment (19). Importantly, SNHG7 has been shown to promote NSCLC development (20,21).
Autophagy is an inducible evolutionarily conserved metabolic process that enables cells to survive under adverse conditions and can be preferentially engaged by tumor cells to enable them to survive under the harsh microenvironmental conditions that they face, including poor nutrient availability, hypoxia, and chemotherapeutic drug exposure (22-24). Several reports have demonstrated a link between autophagy and oncogenic progression owing to its ability to support cancer cell growth, metastasis, and chemoresistance (25). For example, Zhao et al. (26) determined that the lncRNA CRNDE is capable of inducing autophagy in glioblastoma, thereby reducing tumor cell sensitivity to chemotherapeutic treatment. Epigenetic alterations can modify several genes and modulators, eventually leading to inhibition or promotion of autophagy in different cancer stages, and mediating chemoresistance or chemosensitivity (27). Also, lncRNA KCNQ1OT1 reduced small cell lung cancer (SCLC) tumor growth and chemoresistance via the JAK2/STAT3 signaling pathway (28). However, the role of SNHG7 as a regulator of autophagy in DDP-resistant NSCLC has not yet been established, and whether this lncRNA can modulate DDP resistance via the induction of autophagy has yet to be fully clarified. Thus, the present study was developed to explore the role of SNHG7 in DDP-resistant NSCLC and to clarify the underlying mechanisms regulating such activity.
In this study, SNHG7 was found to be upregulated in NSCLC tumors and cells that were DDP-resistant, and its role as a regulator of NSCLC chemoresistance was subsequently explored. We present the following article in accordance with the ARRIVE reporting checklist (available at https://jtd.amegroups.com/article/view/10.21037/jtd-22-1826/rc).
Methods
A protocol was prepared before the study without registration.
Tissue samples
In total, samples were harvested from 105 NSCLC patients undergoing DDP therapy between September 2015 and August 2017 from The Central Hospital of Shaoyang Affiliated to University of South China. During treatment, the patients were assessed to gauge drug efficacy every 2 weeks. NSCLC recurrence was defined as per the Response Evaluation Criteria In Solid Tumors (RECIST) 1.1 criteria for solid tumors as a 20% or 5 mm increase in the total lesion diameter relative to baseline (29,30). NSCLC patient tumors and adjacent paracancerous tissues (>3 cm from malignant tissues) were harvested during surgical biopsy and surgical procedures, including 51 primary DDP-sensitive tissue samples and 54 recurrent DDP-resistant tissue samples, which were immediately stored at −80 ℃ for further analysis.
All patients who participated in this study signed an informed consent form, and this study was approved by the Ethics Committee of The Central Hospital of Shaoyang Affiliated to University of South China (No. KYSQ 2020-026). The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013).
Cell culture
A549 and HCC827 cells (American Type Culture Collection) were cultured in dulbecco minimum essential medium (DMEM, Invitrogen, CA, USA) containing 10% fatal bovine serun (FBS, Invitrogen) and penicillin/streptomycin in a 37 ℃ 5% CO2 incubator. To establish cis-platinum (DDP)-resistant tumor cell lines (A549/DDP and HCC827/DDP), the corresponding parental cell lines were exposed to increasingly high doses of DDP (Sigma, MO, USA) until surviving cells appeared morphologically normal and engaged in expected activities. The resultant A549/DDP and HCC827/DDP cell lines were cultured in standard complete culture media supplemented with DDP (5 µg) to maintain chemoresistance, with DDP being omitted from the media for 1 week prior to experimental use.
Quantitative real-time polymerase chain reaction (qRT-PCR)
Tissue samples from NSCLC patients and the tumors in mice were ground, and the treated A549 and HCC827 cells were collected for qPCR analysis. Briefly, the total RNA was isolated using Trizol (Invitrogen, USA). Then, a Prime Script™ RT Reagent Kit (Takara, Otsu, Japan) and a One-step miRNA RT Kit (Haigene, Harbin, China) were used to prepare complementary DNA (cDNA), after which qPCR assays were performed with a SYBR Green qPCR Mix (Bio-Rad, CA, USA) using the following primers: SNHG7, 5'-ACCTGGTTTGCTCCATGAGG-3' (forward; F), 5'-GTGCCCGAGCTTCAGATACA-3' (reverse; R); Glyceraldehyde-3-phosphate dehydrogenase (GAPDH), 5'-GAGTCAACGGATTTGGTCGT-3' (F), 5'-TTGATTTTGGAGGGATCTCG-3' (R). GAPDH served as a control (31), and the 2−ΔΔCt method was used to assess relative gene expression.
Half maximal inhibitory concentration (IC50) calculation
The IC50 values for DDP were calculated by plating cells in 96-well plates and then replacing the media at 24 h with a media containing a range of DDP concentrations (0, 0.39, 1.56, 3.125, 6.25, 12.5, 25, and 50 µg/mL). After culturing for 48 h, Cell Counting Kit-8 (CCK-8) reagent was added and the cells were cultured for 24 h. Supernatants were then removed and replaced with dimethyl sulfoxide (DMSO), after which absorbance was assessed at 490 nm using a microplate reader (Bio-Rad Laboratories, CA, USA). Experiments were repeated in triplicate.
Cell transfection
An SNHG7-specific shRNA (shSNHG7, 5'-AATTCAAAAAATAATCCGTTTTTACTCCCTCTCTTGAAGTAGCACAACATTCTCCACCCG-3'), a control short hairpin RNA (shRNA) (shCTRL), an SNHG7 overexpression plasmid (OE-SNHG7), and a control empty vector (OE-CTRL) were obtained from Genepharma (Shanghai, China). The cells were transfected using Lipofectamine 2000 (Invitrogen) according to the provided directions. Briefly, the cells were plated in six-well plates for 18 h (5×105/well) until 70–90% confluent and then transfected for 6 h using serum-free DMEM medium (iCell Bioscience Inc., China) containing 100 nM of appropriate siRNA or overexpression constructs. Subsequently, the media was replaced by a standard complete culture media.
Flow cytometry
Harvested NSCLC cells were rinsed using phosphate buffered saline (PBS) and dual stained using a fluorescein isothiocyanate (FITC)-Annexin V apoptosis detection kit (BD Biosciences, NJ, USA), followed by analysis with a flow cytometer (FACSCalibur, BD) to quantify rates of apoptotic death.
Western blotting
Radio-immunoprecipitation assay (RIPA) buffer was used to lyse cells, and the collected protein samples were then separated via sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE) and transferred to polyvinylidene fluoride (PVDF) membranes. Non-specific protein binding was blocked for 1 h via the application of a 5% milk solution, followed by incubation with primary antibodies specific for Beclin1 (1:2,000, ab210498, Abcam, MA, USA), p62 (1:1,500, ab109012, Abcam), and GAPDH (1:10,000, ab9485, Abcam). Blots were then probed with secondary horseradish peroxidase (HRP)-conjugated antibodies (1:20,000, ab205718, Abcam), and protein bands were detected with an enhanced chemiluminescence (ECL) kit (Millipore, Bedford, MA, USA).
Immunofluorescent staining
NSCLC cells were attached to coverslips, fixed using 4% paraformaldehyde, permeabilized using 0.1% Triton X-100, blocked with 3% goat serum, and probed overnight with primary anti-LC3B (Abcam, ab192890, 1:500) at 4 ℃. The samples were then incubated for 1 h with AF488-conjugated anti-rabbit IgG (Invitrogen; 1:500) while protected from light, with the nuclei being counterstained using 4',6-diamidino-2-phenylindole (DAPI, Invitrogen, 1:300).
Terminal deoxynucleotidyltransferase-mediated dUTP-biotin nick end labeling (TUNEL) staining
An in situ cell death detection kit (Beyotime, Beijing, China) was used according to the provided directions for TUNEL staining, with the cells being imaged using a fluorescent microscope (Leica, Cat. #DMI6000B). Apoptotic cells were analyzed based on the relative percentage of TUNEL-positive cells to total cells.
In vivo xenograft modeling and immunohistochemical staining
Nude BALB/c mice (4 weeks old, female) were obtained from Charles River (Beijing, China). A549/DPP cells that had been transfected using shSNHG7 or control constructs were subcutaneously implanted into the right flank of individual mice (5×106/mouse). The mice were additionally administered daily intratumoral injections of PBS, shCTRL, or shSNHG7 combined with intraperitoneal DDP injection (3 mg/kg, every other day) after the tumors were ~100 mm3 in size (32). The tumors were monitored every third day using vernier calipers (each group included six mice). Tumor volumes were measured weekly (volume = length×width2×1/2).
On day 28 post-injection, the mice were euthanized and tumors were excised for qPCR and immunohistochemical (IHC) staining to detect SNHG7 and Ki67 levels, respectively. For IHC staining, anti-Ki-67 (Abcam, ab16667; 1:200) was used, and samples were imaged with a brightfield microscope (Olympus, Tokyo, Japan). The apopsotis cells in tumors were deteted using TUNEL methods. Animal experiments were performed under a project license (No. KYSQ 2020-026) granted by institutional ethics board of The Central Hospital of Shaoyang Affiliated to University of South China, in compliance with institutional guidelines for the care and use of animals.
Statistical analyses
All experiments were performed in triplicate. GraphPad Prism 7.0 (GraphPad Software, CA, USA) and PASW Statistics 18.0 (IBM, SPSS, IL, USA) were used to analyze data, which were compared via one-way analysis of variances (ANOVAs) or Student’s t-tests, with P<0.05 as the significance threshold.
Results
SNHG7 is upregulated in NSCLC and predicts poor patient prognosis
Initially, the expression of SNHG7 was assessed in 105 pairs of NSCLC patient tumors and normal paracancerous tissues via qPCR, which revealed significantly higher SNHG7 levels in tumor samples (Figure 1A). Moreover, samples from DDP-sensitive and DDP-resistant NSCLC patients (n=51 and n=54, respectively) were assessed to explore the link between this lncRNA and chemoresistance. Markedly higher SNHG7 levels were observed in DDP-resistant NSCLC tumors as compared to DDP-sensitive tissues (Figure 1B). After stratifying patients according to the levels of SNHG7 expression, the OS rates were compared between groups using the Kaplan-Meier approach. Increased SNHG7 expression was found to be related to poorer survival among both DDP-sensitive and DDP-resistant patients (Figure 1C,1D). These results suggest that SNHG7 is an essential regulator of NSCLC tumor malignancy and chemoresistance.
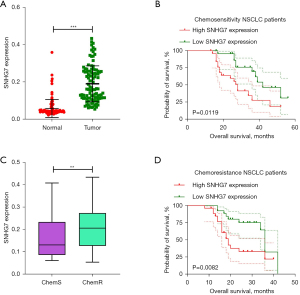
An analysis of the association between SNHG7 expression and the clinicopathological characteristics of these 105 NSCLC patients was additionally conducted (Table 1). This approach revealed higher levels of SNHG7 expression to be closely associated with increased tumor node metastasis (TNM)-stage (P=0.0022), DDP sensitivity (P=0.0012), and lymph node metastasis (P=0.0247). No association was observed between the expression of this lncRNA and patient age, gender, or smoking status. Together, these data indicated that SNHG7 upregulation is a hallmark of NSCLC that is closely related to poorer patient prognosis.
Table 1
| Factors | SNHG7 expression (n=105) | P value | |
|---|---|---|---|
| Low expression (n=52) | High expression (n=53) | ||
| Age, years | 0.4911 | ||
| <60 | 29 | 26 | |
| ≥60 | 23 | 27 | |
| Gender | 0.1438 | ||
| Male | 27 | 20 | |
| Female | 25 | 33 | |
| TNM stage | 0.0022** | ||
| I–II | 37 | 22 | |
| III–IV | 15 | 31 | |
| DDP | 0.0012** | ||
| Sensitive | 36 | 20 | |
| Resistant | 16 | 33 | |
| Lymph node metastasis | 0.0247* | ||
| Yes | 20 | 32 | |
| No | 32 | 21 | |
| Smoking status | 0.6285 | ||
| Smoker | 26 | 29 | |
| Nonsmoker | 26 | 24 | |
*, P<0.05; **, P<0.01. DDP, cisplatin.
SNHG7 is associated with resistance to DDP and the expression of autophagy-related proteins in NSCLC
To examine the functional role of SNHG7 in DDP-resistant NSCLC, two chemoresistant cell lines were generated (A549/DDP and HCC827/DDP), and a CCK-8 assay was used to test the DDP half-maximal IC50 values of these cell lines and the corresponding parental cells as a measure of chemoresistance. As expected, A549/DDP and HCC827/DDP cells exhibited increased IC50 values as compared to parental A549 and HCC827 cells (Figure 2A,2B). Also, marked increases in SNHG7 expression were evident in the DDP-resistant NSCLC cells relative to the corresponding parental cell lines (Figure 2C,2D), suggesting a central role for this lncRNA as a mediator of chemoresistant activity.
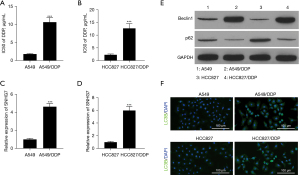
The expression of autophagy-associated proteins including p62 and Beclin1 was then assessed via western blotting in these DDP-resistant and parental NSCLC cell lines (Figure 2E, Figure S1), revealing higher Beclin1 levels and lower p62 levels in the DDP-resistant NSCLC cells. Immunofluorescent staining additionally exhibited significant increases in LC3B expression in A549/DDP and HCC827/DDP cells relative to the corresponding parental controls (Figure 2F). As such, SNHG7 may be linked to autophagy-associated protein levels within NSCLC cells.
SNHG7 regulates NSCLC cell DDP resistance by promoting autophagic and proliferative activity while inhibiting apoptotic cell death
The ability of SNHG7 to modulate chemoresistance, malignancy, autophagy, and apoptosis was next examined in the established NSCLC cell lines by transfecting them with appropriate overexpression (OE-SNHG7) or shRNA (shSNHG7) vectors for gain- and loss-of-function assays. As expected, shSNHG7 transfection successfully suppressed the expression of this lncRNA, whereas its overexpression yielded the opposite effect in the selected DDP-resistant NSCLC cell lines (Figure 3A). Moreover, shSNHG7 transfection reduced the resistance of these cell lines to DDP treatment, while OE-SNHG7 transfection enhanced their ability to resist treatment with this chemotherapeutic drug (Figure 3B). Flow cytometry analyses further revealed that A549/DDP and HCC827/DDP cell apoptosis was markedly enhanced following SNHG7 knockdown, while the opposite outcome was observed following the overexpression of this lncRNA (Figure 3C and Figure S2A). The assessment of autophagy-related protein levels in these cells via western blotting showed that shSNHG7 significantly reduced Beclin1 while promoting p62 upregulation in both DDP-resistant cell lines, while its overexpression yielded the opposite phenotype (Figure 3D and Figure S2B). Similarly, immunofluorescent staining revealed higher LC3B2 levels in DDP-resistant cells in which SNHG7 had been overexpressed as compared to control cells (Figure 3E). Together, these results indicate a role for SNHG7 as a mediator of DDP resistance, functioning in an oncogenic manner to enhance NSCLC cell malignancy.
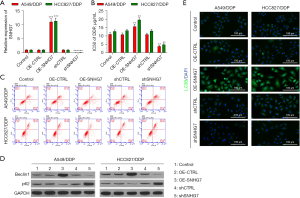
Autophagic induction suppresses the inhibitory impact of SNHG7 knockdown on NSCLC cell chemoresistance
To gain further insight into the interplay between SNHG7 and autophagic induction, the cells were treated with rapamycin (Rapa) to induce autophagy. While SNHG7 knockdown impaired NSCLC cell chemoresistant activity (Figure 3B), Rapa treatment was sufficient to reverse this effect, indicating a role for autophagic induction as a means of counteracting the beneficial effects of knocking down this oncogenic lncRNA in NSCLC (Figure 4A). At 48 h post-transfection, annexin V/PropidiumIodide (PI) flow cytometry analyses revealed significantly lower rates of apoptosis in the shSNHG7 + Rapa group as compared to the shSNHG7 + DMSO and shSNHG7 control groups (Figure 4B,4C), demonstrating a role for SNHG7 knockdown in the induction of apoptotic DDP-resistant NSCLC cell death via the inhibition of autophagy.
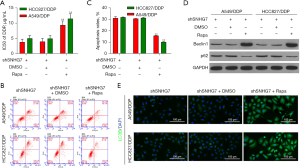
Consistent with these results, Rapa treatment was associated with an increased Baclin1 to LC3B ratio relative to that observed in the shSNHG7 + DMSO and shSNHG7 groups (Figure 4D,4E and Figure S3), confirming that Rapa was able to reverse the chemosensitizing effects of SNHG7 knockdown. Together, these data supported a model in which autophagy can counteract the inhibitory effects of shSNHG7 treatment on NSCLC cell chemoresistance, indicating that reductions in the expression of this lncRNA may lead to the suppression of autophagy, thereby increasing DDP sensitivity in these tumor cells.
SNHG7 knockdown enhances in vivo NSCLC xenograft tumor sensitivity to DDP treatment
A murine xenograft model system was employed to explore the functional role of SNHG7 as a mediator of the chemoresistance of NSCLC cells in vivo. In line with the above in vitro data, the inhibitory antitumor effects of DDP treatment were enhanced in the SNHG7 knockdown group as compared to the blank control and shCTRL groups (Figure 5A). Xenograft tumors derived from shSNHG7-treated A549/DPP cells were significantly smaller than those derived from shCTRL-treated A549/DDP cells at the 4-week stage in mice subjected to DDP treatment (Figure 5B). Importantly, SNHG7 expression was significantly reduced in shSNG7-treated tumor xenografts relative to those from the blank control and shCTRL groups, as assessed via qPCR (Figure 5C).
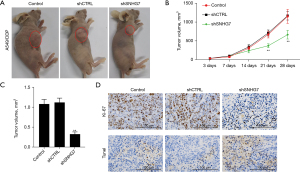
In addition, tumor tissue Ki67 levels were analyzed as a metric of proliferative activity, revealing a reduction in Ki67 expression in the shSNHG7 group as compared to the control and shCTRL groups (Figure 5D and Figure S4). TUNEL staining was employed to analyze tumor cell apoptotic activity, which further revealed that shSNHG7 treatment was associated with an increase in the rates of DDP-induced cell apoptosis (Figure 5D and Figure S4), confirming the ability of this lncRNA to mediate tumor resistance to chemotherapeutic treatment. Together, these results support the above in vitro data and offer clear support for the role of SNHG7 as a novel target for therapeutic efforts aimed at reversing DDP chemoresistance in patients with NSCLC.
Discussion
Despite many recent advances in therapeutic and diagnostic techniques, NSCLC patients continue to face a suboptimal prognosis (3,33). The standard approaches to treating this cancer type include surgery, radiotherapy, chemotherapy, immunotherapy, and specific targeted therapeutic regimens (2). However, the emergence of chemoresistance remains a primary barrier to the effective treatment of patients with this disease. A growing body of evidence has conclusively shown that lncRNAs can influence tumor cell chemosensitivity but the mechanisms underlying such regulatory activity have not yet been firmly established (34,35). Given that the identification of these mechanisms has the potential to shape patient treatment strategies by overcoming therapeutic resistance to NSCLC treatment, further research on this topic is warranted. Herein, SNHG7 upregulation was observed in NSCLC cell lines, and knocking down this lncRNA was sufficient to impair the chemoresistance of DDP-treated HCC827 and A549 cells at least in part by inducing autophagic activity.
SNHG7 is a recently described lncRNA that has been found to be dysregulated in a variety of cancers and associated with an array of oncogenic genes (36), with multiple reports supporting its oncogenic role in lung cancer. For example. She et al. (20) observed SNHG7 upregulation in NSCLC and found that higher levels of this lncRNA correspond to more advanced TNM staging, lymph node metastasis, and poorer patient outcomes. There have also been prior reports linking SNHG7 to chemoresistance, as in the study conducted by Chen et al. (37) who observed the overexpression of this lncRNA in 26 DDP-resistant NSCLC tumors, with loss- and gain-of-function analyses confirming that higher SNHG7 expression levels promote DDP-resistance in these cancer cells. Mechanistically, SNHG7 silencing suppresses p-glycoprotein (MDR1) expression (38), ATP binding cassette subfamily G member 2 (BCRP) expression (39), and PI3K/AKT/mTOR pathway activation (40), all of which are related to chemoresistance. In this report, DDP-resistant NSCLC patient tumor tissues and cell lines exhibited marked SNHG7 upregulation, and the knockdown of this lncRNA confirmed its role as a promoter of NSCLC cell motility, proliferation, and resistance to DDP treatment. Moreover, SNHG7 protected NSCLC cells against apoptotic death.
Autophagy is a tightly regulated and highly conserved metabolic process that regulates cellular homeostatic activity and has been linked to chemoresistance in many cancer types. Indeed, the engagement of autophagic activity can protect tumor cells from chemotherapeutic drugs, improving their survival ability (41,42). For example, Xiong et al. found that overexpressing the hepatocellular carcinoma up-regulated long non-coding RNA (HULC) lncRNA was sufficient to reduce HCC cell sensitivity to chemotherapeutic intervention by upregulating autophagy (43). Similarly, the results of the present study highlight a mechanism whereby SNHG7 can contribute to NSCLC progression and DDP resistance by inducing autophagic activity. DDP-resistant NSCLC tissue samples and cell lines exhibited increased SNHG7 expression as compared to the parental chemosensitive samples, supporting a link between SNHG77 downregulation and the enhanced sensitivity of NSCLC cells to DDP treatment in vitro. Consistently, SNHG7 downregulation enhanced DDP sensitivity, and the induction of autophagy was sufficient to counteract the inhibitory impact of SNHG7 shRNA treatment on NSCLC drug-resistant phenotypes. In addition, in vivo chemosensitivity experiments confirmed that a combination of SNHG7 knockdown and DDP treatment was sufficient to effectively suppress NSCLC tumor growth, consistent with the ability of SNHG7 knockdown to enhance NSCLC cell DDP sensitivity.
Conclusions
Together, these results suggest that the dysregulated expression of SNHG7 strongly impacts NSCLC progression and resistance to DDP treatment, potentially via an autophagy-dependent drug resistance pathway. As such, SNHG7 may offer value as a novel biomarker of poor DDP responsiveness in NSCLC patients and may serve as a promising target for therapeutic intervention in these patients. However, the specific underlying mechanisms linking the regulation of SNHG7 to autophagic activity warrant further investigation.
Acknowledgments
Funding: This study was supported by the Hunan Provincial Health Commission general funding project (No. 202102080487).
Footnote
Reporting Checklist: The authors have completed the ARRIVE reporting checklist. Available at https://jtd.amegroups.com/article/view/10.21037/jtd-22-1826/rc
Data Sharing Statement: Available at https://jtd.amegroups.com/article/view/10.21037/jtd-22-1826/dss
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at https://jtd.amegroups.com/article/view/10.21037/jtd-22-1826/coif). The authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. All patients who participated in this study signed an informed consent form, and this study was approved by the Ethics Committee of The Central Hospital of Shaoyang Affiliated to University of South China (No. KYSQ 2020-026). The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). Animal experiments were performed under a project license (No. KYSQ 2020-026) granted by institutional ethics board of The Central Hospital of Shaoyang Affiliated to University of South China, in compliance with institutional guidelines for the care and use of animals.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Siegel RL, Miller KD, Fuchs HE, et al. Cancer statistics, 2022. CA Cancer J Clin 2022;72:7-33. [Crossref] [PubMed]
- Imyanitov EN, Iyevleva AG, Levchenko EV. Molecular testing and targeted therapy for non-small cell lung cancer: Current status and perspectives. Crit Rev Oncol Hematol 2021;157:103194. [Crossref] [PubMed]
- Tian R, Zhang C, Xiong F, et al. PCAT1/miR-129/ABCB1 axis confers chemoresistance in non-small cell lung cancer. Front Biosci (Landmark Ed) 2020;25:948-60. [Crossref] [PubMed]
- Duma N, Santana-Davila R, Molina JR. Non-Small Cell Lung Cancer: Epidemiology, Screening, Diagnosis, and Treatment. Mayo Clin Proc 2019;94:1623-40. [Crossref] [PubMed]
- Chiang CL, Huang HC, Luo YH, et al. Cerebrospinal fluid as a medium of liquid biopsy in the management of patients with non-small-cell lung cancer having central nervous system metastasis. Front Biosci (Landmark Ed) 2021;26:1679-88. [Crossref] [PubMed]
- Rodríguez-Abreu D, Powell SF, Hochmair MJ, et al. Pemetrexed plus platinum with or without pembrolizumab in patients with previously untreated metastatic nonsquamous NSCLC: protocol-specified final analysis from KEYNOTE-189. Ann Oncol 2021;32:881-95. [Crossref] [PubMed]
- Papadimitrakopoulou VA, Mok TS, Han JY, et al. Osimertinib versus platinum-pemetrexed for patients with EGFR T790M advanced NSCLC and progression on a prior EGFR-tyrosine kinase inhibitor: AURA3 overall survival analysis. Ann Oncol 2020;31:1536-44. [Crossref] [PubMed]
- Cruz-Bermúdez A, Laza-Briviesca R, Vicente-Blanco RJ, et al. Cisplatin resistance involves a metabolic reprogramming through ROS and PGC-1α in NSCLC which can be overcome by OXPHOS inhibition. Free Radic Biol Med 2019;135:167-81. [Crossref] [PubMed]
- Joshi M, Rajender S. Long non-coding RNAs (lncRNAs) in spermatogenesis and male infertility. Reprod Biol Endocrinol 2020;18:103. [Crossref] [PubMed]
- Wang W, Min L, Qiu X, et al. Biological Function of Long Non-coding RNA (LncRNA) Xist. Front Cell Dev Biol 2021;9:645647. [Crossref] [PubMed]
- Luo DB, Lv HB, Sun XH, et al. LncRNA TRERNA1 promotes malignant progression of NSCLC through targeting FOXL1. Eur Rev Med Pharmacol Sci 2020;24:1233-42. [Crossref] [PubMed]
- Chen Z, Chen Q, Cheng Z, et al. Long non-coding RNA CASC9 promotes gefitinib resistance in NSCLC by epigenetic repression of DUSP1. Cell Death Dis 2020;11:858. [Crossref] [PubMed]
- Ge P, Cao L, Zheng M, et al. LncRNA SNHG1 contributes to the cisplatin resistance and progression of NSCLC via miR-330-5p/DCLK1 axis. Exp Mol Pathol 2021;120:104633. [Crossref] [PubMed]
- Ota T, Suzuki Y, Nishikawa T, et al. Complete sequencing and characterization of 21,243 full-length human cDNAs. Nat Genet 2004;36:40-5. [Crossref] [PubMed]
- Bian Z, Ji W, Xu B, et al. The role of long noncoding RNA SNHG7 in human cancers Mol Clin Oncol 2020;13:45. (Review). [Crossref] [PubMed]
- Wu X, Yuan Y, Ma R, et al. lncRNA SNHG7 affects malignant tumor behaviors through downregulation of EZH2 in uveal melanoma cell lines. Oncol Lett 2020;19:1505-15. [Crossref] [PubMed]
- Xia Q, Li J, Yang Z, et al. Long non-coding RNA small nucleolar RNA host gene 7 expression level in prostate cancer tissues predicts the prognosis of patients with prostate cancer. Medicine (Baltimore) 2020;99:e18993. [Crossref] [PubMed]
- Zhao Z, Liu X. LncRNA SNHG7 Regulates Gastric Cancer Progression by miR-485-5p. J Oncol 2021;2021:6147962. [Crossref] [PubMed]
- Jiang C, Qu S, Liu T, et al. Long Noncoding RNA SNHG7 Is a Diagnostic and Prognostic Marker for Colon Adenocarcinoma. Front Oncol 2022;12:893591. [Crossref] [PubMed]
- She K, Yan H, Huang J, et al. miR-193b availability is antagonized by LncRNA-SNHG7 for FAIM2-induced tumour progression in non-small cell lung cancer. Cell Prolif 2018;51:e12406. [Crossref] [PubMed]
- She K, Huang J, Zhou H, et al. lncRNA-SNHG7 promotes the proliferation, migration and invasion and inhibits apoptosis of lung cancer cells by enhancing the FAIM2 expression. Oncol Rep 2016;36:2673-80. [Crossref] [PubMed]
- Hu X, Xuan Y. Bypassing cancer drug resistance by activating multiple death pathways--a proposal from the study of circumventing cancer drug resistance by induction of necroptosis. Cancer Lett 2008;259:127-37. [Crossref] [PubMed]
- Levine B, Klionsky DJ. Development by self-digestion: molecular mechanisms and biological functions of autophagy. Dev Cell 2004;6:463-77. [Crossref] [PubMed]
- Mizushima N, Levine B, Cuervo AM, et al. Autophagy fights disease through cellular self-digestion. Nature 2008;451:1069-75. [Crossref] [PubMed]
- Ferraresi A, Phadngam S, Morani F, et al. Resveratrol inhibits IL-6-induced ovarian cancer cell migration through epigenetic up-regulation of autophagy. Mol Carcinog 2017;56:1164-81. [Crossref] [PubMed]
- Zhao Z, Liu M, Long W, et al. Knockdown lncRNA CRNDE enhances temozolomide chemosensitivity by regulating autophagy in glioblastoma. Cancer Cell Int 2021;21:456. [Crossref] [PubMed]
- Ghavami S, Zamani M, Ahmadi M, et al. Epigenetic regulation of autophagy in gastrointestinal cancers. Biochim Biophys Acta Mol Basis Dis 2022;1868:166512. [Crossref] [PubMed]
- Zhu Y, Shen Y, Chen R, et al. KCNQ1OT1 lncRNA affects the proliferation, apoptosis, and chemoresistance of small cell lung cancer cells via the JAK2/STAT3 axis. Ann Transl Med 2021;9:891. [Crossref] [PubMed]
- Patil T, Mushtaq R, Marsh S, et al. Clinicopathologic Characteristics, Treatment Outcomes, and Acquired Resistance Patterns of Atypical EGFR Mutations and HER2 Alterations in Stage IV Non-Small-Cell Lung Cancer. Clin Lung Cancer 2020;21:e191-204. [Crossref] [PubMed]
- Aras M, Erdil TY, Dane F, et al. Comparison of WHO, RECIST 1.1, EORTC, and PERCIST criteria in the evaluation of treatment response in malignant solid tumors. Nucl Med Commun 2016;37:9-15. [Crossref] [PubMed]
- Yao Z, Xu R, Yuan L, et al. Circ_0001955 facilitates hepatocellular carcinoma (HCC) tumorigenesis by sponging miR-516a-5p to release TRAF6 and MAPK11. Cell Death Dis 2019;10:945. [Crossref] [PubMed]
- Rui Y, Wang D, Hu D, et al. Role of dalteparin sodium on the growth of cancer cells and tumor-associated angiogenesis in A549 human lung cancer cell line and grafted mouse model. J Cancer Res Ther 2018;14:S985-92. [Crossref] [PubMed]
- Evison M. The current treatment landscape in the UK for stage III NSCLC. Br J Cancer 2020;123:3-9. [Crossref] [PubMed]
- Wei L, Sun J, Zhang N, et al. Noncoding RNAs in gastric cancer: implications for drug resistance. Mol Cancer 2020;19:62. [Crossref] [PubMed]
- Wang L, Cho KB, Li Y, et al. Long Noncoding RNA (lncRNA)-Mediated Competing Endogenous RNA Networks Provide Novel Potential Biomarkers and Therapeutic Targets for Colorectal Cancer. Int J Mol Sci 2019;20:5758. [Crossref] [PubMed]
- Zhou Y, Tian B, Tang J, et al. SNHG7: A novel vital oncogenic lncRNA in human cancers. Biomed Pharmacother 2020;124:109921. [Crossref] [PubMed]
- Chen K, Abuduwufuer A, Zhang H, et al. SNHG7 mediates cisplatin-resistance in non-small cell lung cancer by activating PI3K/AKT pathway. Eur Rev Med Pharmacol Sci 2019;23:6935-43. [Crossref] [PubMed]
- Lim JS, Jung GY, Park SY. Nkx-2.5 Regulates MDR1 Expression via Its Upstream Promoter in Breast Cancer Cells. J Korean Med Sci 2019;34:e100. [Crossref] [PubMed]
- Reustle A, Fisel P, Renner O, et al. Characterization of the breast cancer resistance protein (BCRP/ABCG2) in clear cell renal cell carcinoma. Int J Cancer 2018;143:3181-93. [Crossref] [PubMed]
- Lu PW, Li L, Wang F, et al. Inhibitory role of large intergenic noncoding RNA-ROR on tamoxifen resistance in the endocrine therapy of breast cancer by regulating the PI3K/Akt/mTOR signaling pathway. J Cell Physiol 2019;234:1904-12. [Crossref] [PubMed]
- Zhou B, Liu J, Kang R, et al. Ferroptosis is a type of autophagy-dependent cell death. Semin Cancer Biol 2020;66:89-100. [Crossref] [PubMed]
- Kocaturk NM, Akkoc Y, Kig C, et al. Autophagy as a molecular target for cancer treatment. Eur J Pharm Sci 2019;134:116-37. [Crossref] [PubMed]
- Xiong H, Ni Z, He J, et al. LncRNA HULC triggers autophagy via stabilizing Sirt1 and attenuates the chemosensitivity of HCC cells. Oncogene 2017;36:3528-40. [Crossref] [PubMed]
(English Language Editor: A. Kassem)


