
Epithelial-to-mesenchymal transition and drug resistance: transitioning away from death
Cancer metastasis occurs via spreading cancer cells from the primary cancer site to distant organs through the lymphatic system and bloodstream. Lung carcinoma has typical sites of metastasis, such as adrenal glands, bone, brain, liver, and lymph nodes (1). Cancer cell movement from the initial site requires a complicated cellular bioprocess. Epithelial-to-mesenchymal transition (EMT) is a crucial cellular process that promotes metastasis in a variety of tumors by leading to loss of an epithelial cell phenotype, including apical-basal polarity and intracellular junctions, and by gain of mobility through increasing or decreasing a number of genes directed by a small number of EMT transcription factors (2). In addition, as cell phenotype converts from epithelial to mesenchymal, there is almost invariable downregulation of E-cadherin and upregulation of N-cadherin and vimentin (2). Through these changes, cancer cells are freed to migrate, intravasate and eventually invade other tissues, serving as seed cells for metastatic lesions (3).
Several transcription factors, such as the ZEB (ZEB1/2), basic helix loop helix protein 38 (TWIST), and Snail (SNAIL1/2), have been reported as factors that can mediate gene expression (4) and regulate EMT (5). Among these, ZEB1 may be particularly important as a key transcription factor for EMT: in the earliest stages of EMT, ZEB1 is induced by TGF-β signaling, a critical cellular initiator of EMT (6). ZEB1 is correlated with resistance to chemotherapy in cancers and disrupts the epithelial phenotype by repressing epithelial microRNAs like miRNA-200 family members (7). In addition, ZEB1 is broadly involved in EMT in lung, breast, and colorectal cancer through several signaling pathways (8).
While much of the traditional work investigating EMT has been in the context of metastasis, it has also been increasingly appreciated that EMT greatly impacts responses to a broad number of anti-cancer drugs (9-11). This has become increasingly problematic with the growing implementation of targeted therapies, where many secondary resistant mechanisms are druggable, however where EMT is not (12,13) and may in this way possibly be an ultimate and terminal resistant mechanism of a broad number of cancers to a broad number of anti-cancer agents. While some of the resistance associated with EMT involves activation of alternative survival pathways (11,14), how EMT induces resistance to targeted therapies is largely unknown.
To this point, we (4) recently observed that vimentin and ZEB1 are correlated with lower expression of the pro-apoptotic BCL-2 family member, BIM, throughout several cell line and tumor datasets. As BIM is a critical component of the cell death response, in particular following targeted therapies in solid tumors (15-20), we hypothesized that EMT was therefore suppressing apoptosis, which may explain why EMT confers resistance to targeted therapies. By studying EGFR mutant lung cancers treated with EGFR inhibitors (EGFRi), we utilized an EMT system employing a Tet-on/off ER-TWIST plasmid that can activate ZEB1 expression and EMT based on the presence of 4-hydroxytamoxifen (4-OHT). Using this system, we confirmed low expression of BIM and EGFRi efficacy in the EMT-induced cell lines. Mechanistically, we found that ZEB1 bound to and inhibited BIM expression by binding directly to the BIM (BCL2L11) promoter and repressing its expression.
ABT-263 (navitoclax) is a small molecule inhibitor that can induce apoptosis by binding to the anti-apoptotic proteins BCL-2 and BCL-XL, which results in an increase in unbound BIM. We demonstrated that navitoclax is sufficient to restore an apoptotic response in the low BIM expressing EGFRi-resistant EMT cells, resensitizing the cells to EGFRi.
Interestingly, Yochum and colleagues (21) have recently published similar data in EGFR mutant lung cancer. In their study, the authors demonstrate suppression of BIM in EGFR mutant lung cancers that have undergone EMT, which associates with apoptotic resistance and frank resistance. However, instead of focusing on ZEB1, the authors reveal that a fellow EMT transcription factor, TWIST1, can suppress BIM. The authors demonstrate that the navitoclax analog, ABT-737, sensitizes their EMT EGFR mutant lung cancer models to EGFR inhibition. In addition, they identified a TWIST1 inhibitor, harmine, that could also re-sensitize EMT EGFR mutant lung cancers via a similar mechanism of apoptosis sensitization. Together, our studies compliment each other and highlight an understudied and important consequence of EMT—repressed BIM and cell death—that could provide a therapeutic solution to targeting EMT EGFR mutant lung cancers, and potentially other EMT cancers from other targeted therapy paradigms (Figure 1).
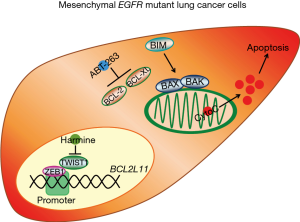
As there is an ongoing clinical phase Ib trial (NCT02520778) combining ABT-263 and AZD 9291 (osimertinib) in previously treated metastatic EGFRi-resistant EGFR mutant non-small cell lung cancer (NSCLC), it would be intriguing if the connection between EMT and response could be elucidated.
So how big of a role does suppressed apoptosis play in EMT-mediated resistance in lung cancer? As mentioned by Marcucci and Rumio (22), EMT may rewire the cell to activate the PI3K pathway in alternative ways (23), and may thus serve as an important component of EMT-mediated drug resistance. This includes activation of other alterative RTKs, like FGFR and AXL (24,25). Along these lines, we recently demonstrated that in KRAS mutant lung cancers, cancers with an EMT phenotype activated PI3K in an ERBB3-independent way, while KRAS mutant lung cancers with an epithelial phenotype activated PI3K in an ERBB3-dependent matter. For the EMT KRAS mutant lung cancers, some were reliant on FGFR1, offering a druggable target (24).
Additionally, as noted by Garinet et al. (26), miRNAs are emerging as another promising target for combination therapy to overcome EMT-mediated drug resistance in lung cancer. MiRNA silencing has been implicated in EMT and tumor invasion and several miRs have been associated with key regulatory pathways, including EGFR and KRAS in NSCLC (27). Forced expression of miR-483-3p, a miRNA significantly silenced in gefitinib-resistant NSCLC cells and lung tissues, increased sensitivity of gefitinib-resistant lung cancer cells to EGFR inhibition by directly targeting integrin β3, thereby repressing downstream FAK/Erk pathway (28).
Therefore, EMT appears to alter the cell in a number of ways, and targeting an apoptotic deficiency, while being an important component, likely will not address all the difficulties EMT cancers present. Given the universal means in which EMT may confer resistance to a large number of anti-cancer drugs in solid tumors, ongoing clinical investigations of navitoclax with different targeted therapies and other anti-cancer agents could yield important information if, for instance, gene expression is measured or histological characterization of the tumors are included.
Acknowledgments
None.
Footnote
Conflicts of Interest: AC Faber has served as a paid consultant for AbbVie. KA Song has no conflicts of interest to declare.
References
- Popper HH. Progression and metastasis of lung cancer. Cancer Metastasis Rev 2016;35:75-91. [Crossref] [PubMed]
- Qi XK, Han HQ, Zhang HJ, et al. OVOL2 links stemness and metastasis via fine-tuning epithelial-mesenchymal transition in nasopharyngeal carcinoma. Theranostics 2018;8:2202-16. [Crossref] [PubMed]
- Deep G, Agarwal R. Antimetastatic efficacy of silibinin: molecular mechanisms and therapeutic potential against cancer. Cancer Metastasis Rev 2010;29:447-63. [Crossref] [PubMed]
- Song KA, Niederst MJ, Lochmann TL, et al. Epithelial-to-Mesenchymal Transition Antagonizes Response to Targeted Therapies in Lung Cancer by Suppressing BIM. Clin Cancer Res 2018;24:197-208. [Crossref] [PubMed]
- Zeisberg M, Neilson EG. Biomarkers for epithelial-mesenchymal transitions. J Clin Invest 2009;119:1429-37. [Crossref] [PubMed]
- Thiery JP, Acloque H, Huang RY, et al. Epithelial-mesenchymal transitions in development and disease. Cell 2009;139:871-90. [Crossref] [PubMed]
- Wellner U, Schubert J, Burk UC, et al. The EMT-activator ZEB1 promotes tumorigenicity by repressing stemness-inhibiting microRNAs. Nat Cell Biol 2009;11:1487-95. [Crossref] [PubMed]
- Graham TR, Yacoub R, Taliaferro-Smith L, et al. Reciprocal regulation of ZEB1 and AR in triple negative breast cancer cells. Breast Cancer Res Treat 2010;123:139-47. [Crossref] [PubMed]
- Oliveras-Ferraros C, Corominas-Faja B, Cufi S, et al. Epithelial-to-mesenchymal transition (EMT) confers primary resistance to trastuzumab (Herceptin). Cell Cycle 2012;11:4020-32. [Crossref] [PubMed]
- Ren J, Chen Y, Song H, et al. Inhibition of ZEB1 reverses EMT and chemoresistance in docetaxel-resistant human lung adenocarcinoma cell line. J Cell Biochem 2013;114:1395-403. [Crossref] [PubMed]
- Wilson C, Nicholes K, Bustos D, et al. Overcoming EMT-associated resistance to anti-cancer drugs via Src/FAK pathway inhibition. Oncotarget 2014;5:7328-41. [Crossref] [PubMed]
- Katayama R, Shaw AT, Khan TM, et al. Mechanisms of acquired crizotinib resistance in ALK-rearranged lung Cancers. Sci Transl Med 2012;4:120ra17. [Crossref] [PubMed]
- Sequist LV, Waltman BA, Dias-Santagata D, et al. Genotypic and histological evolution of lung cancers acquiring resistance to EGFR inhibitors. Sci Transl Med 2011;3:75ra26. [Crossref] [PubMed]
- Wilson C, Ye X, Pham T, et al. AXL inhibition sensitizes mesenchymal cancer cells to antimitotic drugs. Cancer Res 2014;74:5878-90. [Crossref] [PubMed]
- Faber AC, Corcoran RB, Ebi H, et al. BIM expression in treatment-naive cancers predicts responsiveness to kinase inhibitors. Cancer Discov 2011;1:352-65. [Crossref] [PubMed]
- Li Y, Deutzmann A, Choi PS, et al. BIM mediates oncogene inactivation-induced apoptosis in multiple transgenic mouse models of acute lymphoblastic leukemia. Oncotarget 2016;7:26926-34. [PubMed]
- Karachaliou N, Codony-Servat J, Teixido C, et al. BIM and mTOR expression levels predict outcome to erlotinib in EGFR-mutant non-small-cell lung cancer. Sci Rep 2015;5:17499. [Crossref] [PubMed]
- Costa C, Molina MA, Drozdowskyj A, et al. The Impact of EGFR T790M Mutations and BIM mRNA Expression on Outcome in Patients with EGFR-Mutant NSCLC Treated with Erlotinib or Chemotherapy in the Randomized Phase III EURTAC Trial. Clin Cancer Res 2014;20:2001-10. [Crossref] [PubMed]
- Bean GR, Ganesan YT, Dong Y, et al. PUMA and BIM are required for oncogene inactivation-induced apoptosis. Sci Signal 2013;6:ra20. [Crossref] [PubMed]
- Ng KP, Hillmer AM, Chuah CT, et al. A common BIM deletion polymorphism mediates intrinsic resistance and inferior responses to tyrosine kinase inhibitors in cancer. Nat Med 2012;18:521-8. [Crossref] [PubMed]
- Yochum ZA, Cades J, Wang H, et al. Targeting the EMT transcription factor TWIST1 overcomes resistance to EGFR inhibitors in EGFR-mutant non-small-cell lung cancer. Oncogene 2019;38:656-70. [Crossref] [PubMed]
- Marcucci F, Rumio C. From "high" ZEB1 to "low" B-cell lymphoma 2-interacting mediator of cell death (BIM)-an epithelial-mesenchymal transition (EMT)-associated drug resistance pathway elucidated. J Thorac Dis 2019;11:1-5. [Crossref] [PubMed]
- Niederst MJ, Benes CH. EMT twists the road to PI3K. Cancer Discov 2014;4:149-51. [Crossref] [PubMed]
- Kitai H, Ebi H, Tomida S, et al. Epithelial-to-Mesenchymal Transition Defines Feedback Activation of Receptor Tyrosine Kinase Signaling Induced by MEK Inhibition in KRAS-Mutant Lung Cancer. Cancer Discov 2016;6:754-69. [Crossref] [PubMed]
- Zhang Z, Lee JC, Lin L, et al. Activation of the AXL kinase causes resistance to EGFR-targeted therapy in lung cancer. Nat Genet 2012;44:852-60. [Crossref] [PubMed]
- Garinet S, Didelot A, Garelli E, et al. How apoptosis and epithelial-to-mesenchymal transition are nested in EGFR inhibitors resistance in lung cancer. J Thorac Dis 2019;11:47-9. [Crossref] [PubMed]
- Sato H, Shien K, Tomida S, et al. Targeting the miR-200c/LIN28B axis in acquired EGFR-TKI resistance non-small cell lung cancer cells harboring EMT features. Sci Rep 2017;7:40847. [Crossref] [PubMed]
- Yue J, Lv D, Wang C, et al. Epigenetic silencing of miR-483-3p promotes acquired gefitinib resistance and EMT in EGFR-mutant NSCLC by targeting integrin beta3. Oncogene 2018;37:4300-12. [Crossref] [PubMed]


