Performance of an AI based CAD system in solid lung nodule detection on chest phantom radiographs compared to radiology residents and fellow radiologists
Introduction
Chest radiography is an invaluable and cost effective imaging tool used worldwide. In the US alone, 35 million chest radiographs are carried out each year; over 100 chest radiographs are assessed in average by a radiologist a day (1). As an efficient first line diagnostic instrument in various medical conditions, chest radiography has proven its eligibility.
Lung cancer represents a major medical issue and still is a leading cause of cancer-related deaths around the world (2,3). Despite the superior capabilities of computed tomography (CT), chest radiography is still widely used as a first line-imaging tool to screen for and detect lung lesions. However, especially for small lung lesions measuring less than one centimeter, sensitivity of radiographs is low. In addition, the detection of subsolid lesions or ground-glass nodules is significantly inferior as compared to CT (4). Hence, missed lung nodule detection of up to 40% has been reported using chest radiography, resulting in a dissatisfying diagnostic performance regarding the assessment of lung lesions (5,6).
On these grounds, improvement of diagnostic accuracy in reading chest radiographs has been a matter of research in the past, especially computer-aided-diagnostic (CAD) systems have undergone rapid improvements over the last years. Starting in the 1990s with simple tasks such as the correct classification of chest radiograph orientation they have developed to high accuracy definition of the thoracic anatomy, correct lung segmentation or localization of abnormalities such as tuberculosis or lung nodules (7-13).
In recent years, new computer-assisted technologies for radiologic usage, involving deep learning AI systems, have been introduced to clinical research. Deep learning, unlike conventional machine learning, autonomously produces models for presented problems directly from raw data, without the need of human input (14). These deep learning systems showed superior diagnostic performance over conventional CAD systems; one of the first deep learning CAD systems for chest CT showed a sensitivity of 73% and a specificity of 80% in detecting lung nodules, which was considerably better as compared to various conventional CAD systems (15). Recent studies have shown a sensitivity of 85.4% with 1.0 false positive (FP) rate per image (16), or up to 95% however with a FP rate of 1.17 to 22.4 per image (17-19).
Compared to the average diagnostic performance of radiologists with varying experience, deep learning systems showed significantly superior accuracy in distinguishing normal and pathological radiographs as well as detection of lesions, with an overall FP rate per image of 0.11, compared to 0.19 in human readers (20). Other studies have shown lung nodule detection rates in chest radiographs of over 99%, while maintaining a FP rate of 0.2 (21). In reviewing afore by observer only diagnosed radiographs with supplementary deep convolutional network software, reader sensitivity increased from 65.1% to 70.3% and at the same time, the FP rate declined from 0.2 to 0.18 per image (22).
Liang et al. stated that not only the sensitivity of nodule detection increased by the use of an AI algorithm as a second reader device, but that the algorithm could make the daily workflow more efficient (23). The QUIBIM chest X-ray (CXR) classifier achieved rapid processing times of 94.07±16.54 seconds per case (23). Another deep learning based automatic detection algorithm applied in the study of Nam et al. enhanced the performance of both, unexperienced and expert readers, regarding nodule detection and therefore resulted in an optimized workflow as well (24). Besides detecting lung nodules, an AI-CAD system demonstrated benefits in other diagnostic tasks, such as classifying nodules and lung patterns or identifying pulmonary tuberculosis lesions (7,8,25,26).
Despite the undisputed benefits of introducing CAD systems to radiologic practice, there are still challenges requiring further investigation. Conventional machine learning algorithms depend on a large amount of datasets for training purposes; they are therefore limited to the amount and quality of these training datasets. Deep learning algorithms on the other hand are able to gain knowledge and to generalize it to new data. However, they as well require access to well-assorted datasets, which are limited, and depend on inclusion of pathologic verification for most findings (9). Furthermore, most of the recent multicenter studies on this topic dealt with relatively large nodules or even masses, sizes varying between 20–40 mm, resembling locally advanced tumors (20,22).
The primary aim of this study was to test the accuracy of an AI auxiliary nodule detection system on chest phantoms radiographs and compare the performance to radiologists in training, in expectance of a superior performance of the algorithm. The influence of possible confounders such as nodule size, location and experience level of the reader were defined as secondary aims.
We present the following article in accordance with the STARD reporting checklist (available at http://dx.doi.org/10.21037/jtd-20-3522).
Methods
This research project did not involve human data. For this retrospective phantom study, institutional review board approval could be waived.
Chest phantoms
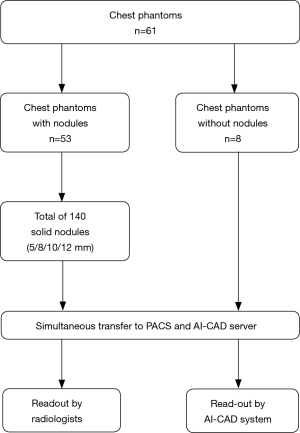
Anthropomorphic chest phantoms (Chest Phantom N1, Kyoto Kagaku) were used to allow for correlation with a robust standard of reference. The phantoms resemble a human male torso containing all the important organs and are made of soft-tissue substitute material (polyurethane) and synthetic bones (epoxy resin), both materials act very similar to the biological tissues regarding their radiation absorption rate. Artificial solid nodules (density: 100 HU; diameters of 5, 8, 10, and 12 mm) were randomly placed inside the phantoms. A random generator chose nodule size, left versus right lung, lung segment and peripheral versus central location for each nodule, the position was documented as a standard of reference.
In total, 61 phantoms were used for this study, 53 phantoms with nodules and 8 without nodules. A total of 140 solid lung nodules were placed in the artificial lung parenchyma (Figure 1). Nodule characteristics are listed in Table 1. The chest radiograph unit settings for the phantoms were identical to those used in the clinical setting. Radiographs were acquired in two planes in upright position, including dual-energy subtraction of the bones. This experimental design was replicated from an earlier study, which dealt with the detection rate of radiologists in chest radiographies compared to chest CT scans with varying kernels (27).
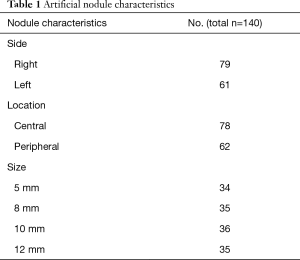
Full table
AI readout
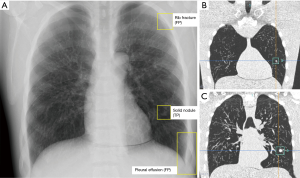
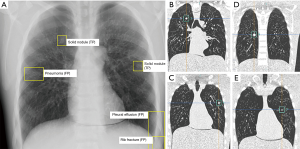
The phantom scans were transferred to an in-house server running a commercially available machine-learning algorithm for lung lesion detection on chest radiographs (InferRead® DR by Infervision Technology Ltd., Beijing, China). The algorithm is based on a region-based fully convolutional network (R-FCN network), which is a region-based full convolution detection method, consisting of several sub-network structures. The first network performs convolutional operation on the entire image to obtain the corresponding feature map, while the second is the region proposal network (RPN) used to generate the candidate area, and the third is the sub-network used for classification and regression of the position of the detection frame, which is regionally correlated (28). Prior to the deployment, the algorithm was tuned to the site-specific characteristics of the radiography unit. The dataset used for adjustment comprised 20 radiographs, which were retrospectively retrieved from the clinical database. The cases were irreversibly anonymized and transferred to Infervision for algorithm adaption. After completion of the installation, the test dataset of phantom scans was transferred to the AI-CAD. Automated analysis of each case was performed (Figures 2 and 3) and the results were recorded by a chest radiology fellow on a spread sheet. Results were compared to the ground truth. Correctly identified lesions were recorded as true positives (TP), missed lesions as false negatives (FN), correctly identified healthy controls as true negatives (TN) and incorrectly identified lesions as FP.
Human readout
Seven radiologists (two fellows in thoracic imaging with 5 years of experience and five residents with 2–4 years of experience in general radiology, including at least 1 year of experience in thoracic imaging) independently conducted a blinded retrospective readout. For the readout of the conventional chest radiographs, the radiologists evaluated the radiographs in both planes and subsequently interpreted the posterior-anterior projection with bone subtraction. The readout was performed on a dedicated PACS-workstation (Sectra PACS IDS7, Sectra) with dedicated monitors (BARCO Coronis Fusion 6MP LED). The readers were allowed to adjust window settings to allow for natural reading conditions. The readers, who were blinded to the nodule distribution, searched the radiographs for lung nodules. Lesions were recorded on a standardized readout sheet by documenting their location. Results were compared to the ground truth, analogous to the AI readout.
Statistical analysis
All data analysis was performed with Microsoft Excel (Microsoft Excel, Version 2016, Redmond, WA, USA) and SPSS (IBM SPSS Statistics., Version 25.0. Armonk, NY, USA).
Sensitivity, specificity and accuracy of the software and each reader were calculated and McNemar test was used to compare each radiologist’s results to the software. A P value of <0.05 was considered statistically significant. Additionally, weighted Fleiss-Kappa was applied for the inter-rater and intra-observer agreements.
Results
Per nodule analysis
The AI-CAD software showed a sensitivity of 0.35 (P=0.001), specificity of 0.84 (P=0.79) and accuracy of 0.47 (P=0.02) in nodule detection.
In the individual performance comparison between each reader and the AI-CAD, every radiologist’s accuracy was significantly higher than the AI-CAD software, except for residents 4 and 5, who demonstrated statistically the same accuracy of 0.56 (P=0.085) and 0.55 (P=0.122), respectively (Table 2).
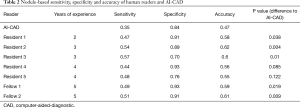
Full table
The readers reached a pooled sensitivity, specificity and accuracy of 0.50, 0.86 and 0.59. An experience dependent evaluation was performed; sensitivity, specificity and accuracy of the residents were 0.50, 0.84 and 0.58. Sensitivity, specificity and accuracy of the fellows were 0.50, 0.92 and 0.60, respectively. Sensitivity and accuracy did not demonstrate a significant difference between residents and fellows (P values: 0.92 and 0.59) while the specificity of the fellows was significantly higher (P value =0.046). Compared to the AI-CAD software, sensitivity and accuracy of both residents and fellows were significantly better (P<0.005) and there was no difference in specificity (residents vs. AI-CAD: P=0.87 and fellows vs. AI-CAD: P=0.17).
Inter-rater agreement among the radiologists was higher (K’=0.58±0.13) than the agreement between radiologists and AI-CAD software (K’=0.51±0.1; P<0.001).
Additionally, intra-observer agreements for resident 2 and fellow 1 were calculated and showed substantial to almost perfect agreement in a second reading session, which was performed 6 months after the initial reading session to avoid recall bias (resident 2: K’=0.83±0.02; fellow 1: K’=0.78±0.02).
Per phantom analysis
A phantom-based evaluation of the results was performed, meaning that for a TP read at least one nodule had to be correctly identified or a normal control had to be read as normal.
Similar to the nodule based approach all radiologists except residents 4 and 5 showed significantly higher accuracy than the AI-CAD software (Table 3). The radiologists reached a pooled sensitivity, specificity and accuracy of 0.83, 0.80 and 0.82 respectively, while the AI-CAD software reached 0.67, 0.66 and 0.75. The differences in accuracy and sensitivity between the software and the radiologists were significant (P=0.005 and P=0.004 respectively) while specificity was not significantly different (P=0.72). The experience dependent evaluation showed no significant differences between fellows and residents regarding accuracy (P=0.49), sensitivity (P=0.93) or specificity (P=0.11). Inter-rater agreements between the radiologists (K’=0.61±0.16, substantial) were higher than the agreement between radiologists and software (K’=0.43±0.09, moderate).
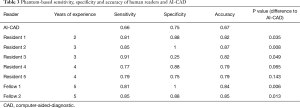
Full table
Intra-observer agreements were calculated for resident 2 and fellow 1 on the per phantom basis as well and each of them showed almost perfect agreement (resident 2: K’=0.85±.05; fellow 1: K’=0.80±.02) in the second reading session 6 months after the initial session.
Effect of nodule size and location
Radiologists’ average sensitivity for 12 and 10 mm nodules was 0.76 and 0.82 respectively and dropped to 0.66 for the nodule size of 8 mm (P=0.04). For the 5 mm nodules the sensitivity reached a minimum of 0.14 (P value <0.001, compared to the 8 mm nodules). The results were similar amongst fellows, residents and the AI-CAD; however, there was no significant effect of nodule size on the results of the AI-CAD.
Both, the radiologists in training and the AI-CAD demonstrated a significant higher sensitivity for peripheral nodules (outer 2/3 of the lung on axial CT images) compared to central nodules (radiologists: 0.66 vs. 0.47, P=0.004; AI-CAD: 0.64 vs. 0.09, P=0.025).
A size dependent analysis of nodule detection sensitivity showed, that radiologists in general detected significantly more nodules as compared to the AI-CAD (P=0.009). However, only in lesions measuring 10 mm in diameter, results were found to be significant (P=0.005; Table 4).

Full table
Discussion
The purpose of this study was to evaluate the performance of a commercially available AI based CAD system regarding the detection of solid lung nodules on chest radiograph phantoms. The results of the software were compared to seven radiologists in training; five of them achieved a significant higher accuracy than the AI-based CAD system.
The software showed a sensitivity of 0.35 regarding the detection of lung nodules, which is lower compared to the sensitivities reached by comparable algorithms in recent studies, for example, the deep convolutional neural network (DCNN) software in the study of Sim et al. (0.67) or a deep learning-based detection (DLD) system in the study of Park and colleagues (0.84) (20,22). The most probable reason is the use of chest phantoms instead of real patients; the system had initially been trained with a dataset of human chest radiographs. A hint in this direction is that the system detected many false-positive ancillary findings, which were not present in the phantoms (e.g., pleural effusion, rib fractures, pneumonia). However, additional studies with human datasets are needed in the future to confirm this theory.
Another reason for the system’s performance most certainly was the use of micronodules and small nodules in the current study (5, 8, 10, 12 mm). This is pertinent, keeping in mind that the formerly mentioned multicenter studies all included much larger nodules or even masses; for example Sim et al. included nodules between 10–30 mm while Park et al. had average nodule sizes of 41±26 mm in center 1 and 34±18 mm in center 2 (20,22).
In the current study, an additional phantom-based evaluation was performed, meaning that with one TP nodule or a correctly identified healthy control the phantom was evaluated as correctly read. Based on this per-phantom approach, the system’s sensitivity was 0.66; but again human readers achieved higher sensitivities and accuracies.
According to the results of the current study, the location of the nodules was more important than their size in terms of detection. Both, radiologists and algorithm, detected more peripheral than central nodules, which is an expected finding. Still, there was only moderate inter-rater agreement between algorithm and the human readers, a finding that may indicate that the AI-CAD system detected different nodules than the radiologists and therefore the two may work complementary. Of course, based on the current results this can be only hypothesized, but it is in agreement with previous studies, of which most concluded that CAD systems should rather be used as a second reader than as a concurrent or primary reader in the context of CT lung cancer screening (29-31).
Most of the deep learning algorithms act inferior if compared to the human learning capability, which as well is in concordance with the findings of the current study (32). One also has to keep in mind that most algorithms are tested on datasets, which are very similar to the training datasets, a fact that may cause an overestimation of their performance; this phenomenon has been described in the literature as “overfitting” (33). Despite the promising results of AI-supported diagnostic modules, widespread implementation is still delayed; mostly owed to inadequate performance, lack of workflow integration and assessment tools (32,33).
Another important aspect for radiography interpretation by AI algorithms is the identification of ancillary findings, of which most seem trivial for the human radiologist, but CAD systems need to be taught first. In the current study, the AI-CAD system revealed difficulties with ancillary findings and had a high number of FP findings such as pneumonia, rib fractures or atelectasis, which were not present in the phantoms. This most probably was caused by the use of chest phantoms instead of real patients; it may be diminished by further optimization of the algorithm and has to be confirmed by future studies on human datasets.
Further reported pertinent problems for the implementation of AI based systems include limits of generalization, the lack of explainability and a lack of publication standards for reproducibility (15).
Despite the decreasing relevance of chest radiography in the field of lung cancer screening, there is still a good chance that the diagnostic work-up of a symptomatic patient starts with a chest radiography to get a first impression of his pulmonary status or to rule out other relevant differentials such as pneumonia or pneumothorax. Even though there is decreased sensitivity for small nodules and subsolid or ground-glass nodules documented in the literature, radiographs are still a pivotal tool for primary assessment of the lungs (4,34,35).
There are several limitations to the current study, the first one being the use of chest phantoms instead of human patients, which most probably lead to a rather poor performance of the algorithm but on the other hand allowed for correlation with a robust ground truth. Another limitation is that only solid nodules were used and no part-solid or ground-glass nodules were available for this study. However, it is questionable if those nodules are detectable with radiography at all. Finally, the small sample size could be considered a limitation, especially for the calculation of the inter-rater agreements, but the achieved results proved to be mainly statistically significant.
Conclusions
In conclusion, we found that human readers show superior accuracy as compared to an AI based CAD system. However, the AI-CAD system used did detect different lesions than the radiologists; emphasizing the role of such a system as a second reader device.
Acknowledgments
Funding: None.
Footnote
Reporting Checklist: The authors have completed the STARD reporting checklist. Available at http://dx.doi.org/10.21037/jtd-20-3522
Data Sharing Statement: Available at http://dx.doi.org/10.21037/jtd-20-3522
Peer Review File: Available at http://dx.doi.org/10.21037/jtd-20-3522
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jtd-20-3522). Dr. JHT reports grants from Bayer Healthcare AG, grants from Guerbet AG, grants from Siemens Healthineers and grants from Bracco Imaging Spa, outside the submitted work. The other authors have no conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. This research project did not involve human data. For this retrospective phantom study, institutional review board approval could be waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Kamel SI, Levin DC, Parker L, et al. Utilization trends in noncardiac thoracic imaging, 2002-2014. J Am Coll Radiol 2017;14:337-42. [Crossref] [PubMed]
- Torre LA, Bray F, Siegel RL, et al. Global cancer statistics, 2012. CA Cancer J Clin 2015;65:87-108. [Crossref] [PubMed]
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin 2020;70:7-30. [Crossref] [PubMed]
- Detterbeck FC, Homer RJ. Approach to the ground-glass nodule. Clin Chest Med 2011;32:799-810. [Crossref] [PubMed]
- Finigan JH, Kern JA. Lung cancer screening: Past, Present and Future. Clin Chest Med 2013;34:365-71. [Crossref] [PubMed]
- Quekel LG, Kessels AG, Goei R, et al. Miss rate of lung cancer on the chest radiograph in clinical practice. Chest 1999;115:720-4. [Crossref] [PubMed]
- Kallianos K, Mongan J, Antani S, et al. How far have we come? Artificial intelligence for chest radiograph interpretation. Clin Radiol 2019;74:338-45. [Crossref] [PubMed]
- Qin C, Yao D, Shi Y, et al. Computer-aided detection in chest radiography based on artificial intelligence: A survey. BioMed Eng OnLine 2018;17:113. [Crossref] [PubMed]
- Cha MJ, Chung MJ, Lee JH, et al. Performance of Deep Learning Model in Detecting Operable Lung Cancer With Chest Radiographs. J Thorac Imaging 2019;34:86-91. [Crossref] [PubMed]
- Boone JM, Seshagiri S, Steiner RM. Recognition of chest radiograph orientation for picture archiving and communications systems display using neural networks. J Digit Imaging 1992;5:190-3. [Crossref] [PubMed]
- Sivaramakrishnan R, Antani S, Xue Z, et al. Visualizing abnormalities in chest radiographs through salient network activations in Deep Learning. 2017 IEEE Life Sciences Conference (LSC), Sydney, NSW, 2017;71-4.
- Jaeger S, Karargyris A, Candemir S, et al. Automatic tuberculosis screening using chest radiographs. IEEE Trans Med Imaging 2014;33:233-45. [Crossref] [PubMed]
- Xue Z, Jaeger S, Antani S, et al. Localizing tuberculosis in chest radiographs with deep learning. Medical imaging 2018: imaging informatics for healthcare, research, and applications. International Society for Optics and Photonics 2018;105790U.
- Lee SM, Seo JB, Yun J, et al. Deep Learning Applications in Chest Radiography and Computed Tomography: Current State of the Art. J Thorac Imaging 2019;34:75-85. [Crossref] [PubMed]
- Hua KL, Hsu CH, Hidayati SC, et al. Computer-aided classification of lung nodules on computed tomography images via deep learning technique. Onco Targets Ther 2015;8:2015-22. [PubMed]
- Setio AAA, Ciompi F, Litjens G, et al. Pulmonary nodule detection in CT images: false positive reduction using multi-view convolutional networks. IEEE Trans Med Imaging 2016;35:1160-9. [Crossref] [PubMed]
- Hamidian S, Sahiner B, Petrick N, et al. 3D Convolutional neural network for automatic detection of lung nodules in chest CT. Proc SPIE Int Soc Opt Eng 2017;10134:1013409.
- Masood A, Sheng B, Li P, et al. Computer-assisted decision support system in pulmonary cancer detection and stage classification on CT images. J Biomed Inform 2018;79:117-28. [Crossref] [PubMed]
- Jiang H, Ma H, Qian W, et al. An Automatic Detection System of Lung Nodule Based on Multigroup Patch-Based Deep Learning Network. IEEE J Biomed Health Inform 2018;22:1227-37. [Crossref] [PubMed]
- Park S, Lee SM, Lee KH, et al. Deep learning-based detection system for multiclass lesions on chest radiographs: comparison with observer readings. Eur Radiol 2020;30:1359-68. [Crossref] [PubMed]
- Li X, Shen L, Xie X, et al. Multi-resolution convolutional networks for chest X-ray radiograph based lung nodule detection. Artif Intell Med 2020;103:101744 [Crossref] [PubMed]
- Sim Y, Chung MJ, Kotter E, et al. Deep Convolutional Neural Network-based Software Improves Radiologist Detection of Malignant Lung Nodules on Chest Radiographs. Radiology 2020;294:199-209. [Crossref] [PubMed]
- Liang CH, Liu YC, Wu MT, et al. Identifying pulmonary nodules or masses on chest radiography using deep learning: external validation and strategies to improve clinical practice. Clin Radiol 2020;75:38-45. [Crossref] [PubMed]
- Nam JG, Park S, Hwang EJ, et al. Development and Validation of Deep Learning-based Automatic Detection Algorithm for Malignant Pulmonary Nodules on Chest Radiographs. Radiology 2019;290:218-28. [Crossref] [PubMed]
- Lakhani P, Sundaram B. Deep learning at chest radiography: Automated Classification of Pulmonary Tuberculosis by Using Convolutional Neural Networks. Radiology 2017;284:574-82. [Crossref] [PubMed]
- Christodoulidis S, Anthimopoulos M, Ebner L, et al. Lung Pattern Classification for Interstitial Lung Diseases Using a Deep Convolutional Neural Network. IEEE J Biomed Health Inform 2017;21:76-84. [Crossref] [PubMed]
- Ebner L, Bütikofer Y, Ott D, et al. Lung Nodule Detection by Microdose CT Versus Chest Radiography (Standard and Dual-Energy Subtracted). AJR 2015;204:727-35. [Crossref] [PubMed]
- Dai J, Li Y, He K, et al. R-FCN: Object Detection via Region-based Fully Convolutional Networks. Neural Information Processing Systems 2016:379-87.
- Sahiner B, Chan HP, Hadjiiski LM, et al. Effect of CAD on radiologists’ detection of lung nodules on thoracic CT scans: analysis of an observer performance study by nodule size. Acad Radiol 2009;16:1518-30. [Crossref] [PubMed]
- Beigelman-Aubry C, Raffy P, Yang W, et al. Computer-aided detection of solid lung nodules on follow-up MDCT screening: evaluation of detection, tracking, and reading time. AJR 2007;189:948-55. [Crossref] [PubMed]
- Liang M, Tang W, Xu DM, et al. Low-Dose CT Screening for Lung Cancer: Computer-aided Detection of Missed Lung Cancers. Radiology 2016;281:279-88. [Crossref] [PubMed]
- van Ginneken B, Schaefer-Prokop CM, Prokop M. Computer-aided diagnosis: how to move from the laboratory to the clinic. Radiology 2011;261:719-32. [Crossref] [PubMed]
- Tandon YK, Bartholmai BJ, Koo CW. Putting artificial intelligence (AI) on the spot: machine learning evaluation of pulmonary nodules. J Thorac Dis 2020;12:6954-65. [Crossref] [PubMed]
- Kruger R, Flynn MJ, Judy PF, et al. Effective dose assessment for participants in the National Lung Screening Trial undergoing posteroanterior chest radiographic examinations. AJR 2013;201:142-6. [Crossref] [PubMed]
- Shah PK, Austin JH, White CS, et al. Missed nonsmall cell lung cancers: radiographic findings of potentially resectable lesions evident only in retrospect. Radiology 2003;226:235-41. [Crossref] [PubMed]