Towards engineering integrated cardiac organoids: beating recorded
Introduction
Modeling human physiology in vitro is burgeoning as an exciting new field that requires coherent efforts encompassing multiple disciplines including engineering, biology, and medicine, among others (1). Originating from the concept of tissue engineering (2), yet with a distinct aim that is to construct models of human tissues and organs outside the body for improved biological, pharmaceutical, and environmental studies (3-6) rather than repairing them in vivo, these microphysiological systems have further undergone significant developments with the inclusion of the microfabrication and microfluidics technologies that conveniently bring in the beneficial complexity (4,7). While various microfabrication strategies allow us to engineer microscale tissue and organ units that possess shapes and architecture that mimic their in vivo counterparts (8-10), the ability to manipulate fluids at small scales leads to reproduction of the dynamic microenvironments indispensable for the functions of natural tissues and organs (1,4), both of which are otherwise not achievable using the conventional planar, static culture platforms. Of note, these individual microphysiological systems can be further linked together in such a way that the interconnected multi-unit platforms recapitulate the linkage of the various tissues and organs in their native arrangements, facilitating investigations of the intricate interactions among these different components in vitro (3-6).
To date, a variety of microphysiological systems have been developed that model their respective tissues and organs (and their disease states) of the human body, ranging from those mimicking the nervous system (11), the respiratory system (12,13), the digestive system (14,15), and the musculoskeletal system (16,17) to the cardiovascular system (18,19), covering essentially every single type of tissue and organ (4,5). Among all, the cardiac organoids have attracted increasing attention due to their critical roles in toxicology; for example, it is estimated that cardiotoxicity represents a major side effect of systemic drug toxicity—in the past 40 years 19% of drug recalls were likely due to cardiotoxicity (20).
A human heart contains four main chambers of two atria and two ventricles as well as valves and heart wall laden with specialized cell populations. The myocardium is responsible for contraction of the heart, which is primarily composed of cardiomyocytes densely aligned in parallel forming the bundled myocardial fibers and enhancing the contractile force to pump the blood throughout the vascular system of the body (21). Besides cardiomyocytes, approximately half of the cells in the heart are cardiac fibroblasts that produce the connective and elastic extracellular matrix (ECM) of the heart wall (21). Other important cell populations include the cardiac signal conduction system made of pacemakers cells and Purkinje fibers (21), as well as the endothelial cells that form the vasculature and supply nutrients to the cardiac cells (18).
Various approaches have been exploited to fabricate cardiac organoids and heart-on-a-chip devices that recapitulate the biology and physiology of a native heart. For instance, microengineering strategies that rely on the use of topographic cues would induce alignment of the cardiomyocytes that assume a similar structure of the native myocardium (22); bioprinting strategies have enabled the incorporation of vasculature into the cardiac organoids while maintaining the anisotropy of the cardiomyocytes (23); and the adoption of flexible substrates seeded with cardiomyocytes could lead to spontaneously actuating heart wall-like patterns modeling a part of a beating heart (24).
While significant efforts have been exerted on engineering functional cardiac organoids, ways to monitor their behaviors and responses have been limited. Common sensing strategies mainly rely on optical methods, either by imaging the contraction of the cardiac organoids (24,25), or by mapping the actions of the cardiomyocytes [e.g., Ca2+ flux following staining with fluorescence dyes (26)]. Besides, electrophysiological signals could also be measured by depositing microelectrode arrays onto the substrate on which the cardiomyocytes are housed (27). Nevertheless, these sensing elements have rarely been able to provide accurate, conformal measurements of the cardiac organoids as they typically would only represent projected collective behaviors (in the case of optical mapping) or planar signals (in the case of electrophysiology) of the model systems.
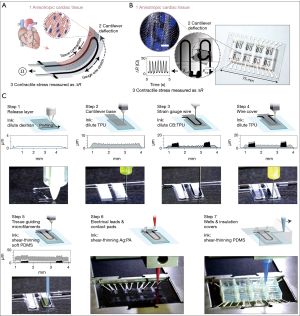
A recent publication by Lind et al., shed light on the existing dilemma, which for the first time provided an enabling strategy to fabricate fully integrated sensor-impregnated heart-on-a-chip device (Figure 1A,B) (28). Specifically, they used a three-dimensional (3D) multi-material extrusion printer with a multi-step procedure to generate the thin-film cantilever capable of deflecting by seeded cardiomyocytes, whereas the embedded conductive wires could directly measure the contractile signals during the deflections that would reflect the status of the cardiac organoids (Figure 1C): (I) printing the 0.5 µm-thick bottom release layer of dextran; (II) printing the 3 µm-thick cantilever base of thermoplastic polyurethane (TPU); (III) printing the 6.5 µm-thick strain gauge wires of TPU encapsulating 25 wt.% conductive carbon black nanoparticles; (IV) printing the 1.5 µm-thick TPU wire cover; (V) printing cardiomyocyte-guiding microfilaments of polydimethylsiloxane (PDMS); (VI) printing electrical leads and contact pads of silver:polyamide (Ag:PA); (VII) printing the well structure and insulation covers of PDMS; and (VIII) seeding the cardiomyocytes onto the printed cantilevers following curing of all the materials.
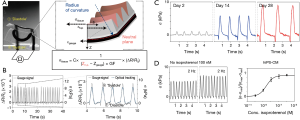
Applying such a heart-on-a-chip system, the embedded strain gauge could conveniently measure the signals generated by each contracting cantilever via establishing a mechanical model that provided conversion between the changes in gauge resistance and those in cantilever radius/curvature, expressed as tissue twitch stress (σ), i.e., the difference between the systolic and diastolic stresses (Figure 2A). The contracting signals measured with the printed gauge on the cantilever were found to well correlate with the conventional optical tracking (Figure 2B), indicating the accuracy of the integrated sensing units and eliminating the needs for external instrumentation such as the use of a dedicated microscope. This sensor-impregnated heart-on-a-chip platform not only allowed for long-term culture and maturation of the human induced pluripotent stem cell-derived cardiomyocytes (hiPS-CMs, Figure 2C) but more importantly, the real-time, in-line recording of their dose-dependent responses to pharmaceutical compounds (Figure 2D).
This piece of work represents the first example of constructing a flexible organ-on-a-chip platform that is directly impregnated with sensing units for signal recording. The integration of the sensing elements is instrumental, allowing for seamless data collection and readouts of organoid status with no need of secondary devices. The conformal nature of the sensors further resulted in more accurate measurements that are less affected by the spatial movement of the 3D organoids than their projection-based or planar counterparts where the signals might be biased. Although currently the technology is still in its infant stage with limited complexity in the circuitry and types of signals that can be measured, its further optimization, development, and expansion into the combination with more sophisticated organoids potentially even diseased models [e.g., mitochondrial cardiomyopathy (29)], are envisioned in the near future.
Acknowledgements
Funding: YS Zhang acknowledges the National Cancer Institute of the National Institutes of Health Pathway to Independence Award (K99CA201603). C Yu acknowledges the financial support from University of Houston (Award for Excellence in Research, Scholarship or Creative Activity) and National Science Foundation (NSF-CMMI-1554499).
Footnote
Conflicts of Interest: The authors have no conflicts of interest to declare.
References
- Zhang YS, Khademhosseini A. Seeking the right context for evaluating nanomedicine: from tissue models in petri dishes to microfluidic organs-on-a-chip. Nanomedicine (Lond) 2015;10:685-8. [Crossref] [PubMed]
- Langer R, Vacanti JP. Tissue engineering. Science 1993;260:920-6. [Crossref] [PubMed]
- Esch MB, King TL, Shuler ML. The role of body-on-a-chip devices in drug and toxicity studies. Annu Rev Biomed Eng 2011;13:55-72. [Crossref] [PubMed]
- Bhatia SN, Ingber DE. Microfluidic organs-on-chips. Nat Biotechnol 2014;32:760-72. [Crossref] [PubMed]
- Wikswo JP. The relevance and potential roles of microphysiological systems in biology and medicine. Exp Biol Med (Maywood) 2014;239:1061-72. [Crossref] [PubMed]
- Skardal A, Shupe T, Atala A. Organoid-on-a-chip and body-on-a-chip systems for drug screening and disease modeling. Drug Discov Today 2016;21:1399-411. [Crossref] [PubMed]
- Ingber DE. Reverse Engineering Human Pathophysiology with Organs-on-Chips. Cell 2016;164:1105-9. [Crossref] [PubMed]
- Khademhosseini A, Langer R. Microengineered hydrogels for tissue engineering. Biomaterials 2007;28:5087-92. [Crossref] [PubMed]
- Zhang YS, Yue K, Aleman J, et al. 3D Bioprinting for Tissue and Organ Fabrication. Ann Biomed Eng 2016. [Epub ahead of print].
- Zhang YS, Xia Y. Multiple facets for extracellular matrix mimicking in regenerative medicine. Nanomedicine (Lond) 2015;10:689-92. [Crossref] [PubMed]
- Yi Y, Park J, Lim J, et al. Central Nervous System and its Disease Models on a Chip. Trends Biotechnol 2015;33:762-76. [Crossref] [PubMed]
- Huh D, Matthews BD, Mammoto A, et al. Reconstituting organ-level lung functions on a chip. Science 2010;328:1662-8. [Crossref] [PubMed]
- Benam KH, Villenave R, Lucchesi C, et al. Small airway-on-a-chip enables analysis of human lung inflammation and drug responses in vitro. Nat Methods 2016;13:151-7. [Crossref] [PubMed]
- Kim HJ, Li H, Collins JJ, et al. Contributions of microbiome and mechanical deformation to intestinal bacterial overgrowth and inflammation in a human gut-on-a-chip. Proc Natl Acad Sci U S A 2016;113:E7-15. [Crossref] [PubMed]
- Bale SS, Vernetti L, Senutovitch N, et al. In vitro platforms for evaluating liver toxicity. Exp Biol Med (Maywood) 2014;239:1180-91. [Crossref] [PubMed]
- Fong EL, Lamhamedi-Cherradi SE, Burdett E, et al. Modeling Ewing sarcoma tumors in vitro with 3D scaffolds. Proc Natl Acad Sci U S A 2013;110:6500-5. [Crossref] [PubMed]
- Grosberg A, Nesmith AP, Goss JA, et al. Muscle on a chip: in vitro contractility assays for smooth and striated muscle. J Pharmacol Toxicol Methods 2012;65:126-35. [Crossref] [PubMed]
- Zhang YS, Aleman J, Arneri A, et al. From cardiac tissue engineering to heart-on-a-chip: beating challenges. Biomed Mater 2015;10:034006. [Crossref] [PubMed]
- Wong KH, Chan JM, Kamm RD, et al. Microfluidic models of vascular functions. Annu Rev Biomed Eng 2012;14:205-30. [Crossref] [PubMed]
- Piccini JP, Whellan DJ, Berridge BR, et al. Current challenges in the evaluation of cardiac safety during drug development: translational medicine meets the Critical Path Initiative. Am Heart J 2009;158:317-26. [Crossref] [PubMed]
- Olson EN. Gene regulatory networks in the evolution and development of the heart. Science 2006;313:1922-7. [Crossref] [PubMed]
- Schroer AK, Shotwell MS, Sidorov VY, et al. I-Wire Heart-on-a-Chip II: Biomechanical analysis of contractile, three-dimensional cardiomyocyte tissue constructs. Acta Biomater 2016. [Epub ahead of print]. [Crossref] [PubMed]
- Zhang YS, Arneri A, Bersini S, et al. Bioprinting 3D microfibrous scaffolds for engineering endothelialized myocardium and heart-on-a-chip. Biomaterials 2016;110:45-59. [Crossref] [PubMed]
- Agarwal A, Goss JA, Cho A, et al. Microfluidic heart on a chip for higher throughput pharmacological studies. Lab Chip 2013;13:3599-608. [Crossref] [PubMed]
- Zhang YS, Ribas J, Nadhman A, et al. A cost-effective fluorescence mini-microscope for biomedical applications. Lab Chip 2015;15:3661-9. [Crossref] [PubMed]
- McCain ML, Sheehy SP, Grosberg A, et al. Recapitulating maladaptive, multiscale remodeling of failing myocardium on a chip. Proc Natl Acad Sci U S A 2013;110:9770-5. [Crossref] [PubMed]
- Kujala VJ, Pasqualini FS, Goss JA, et al. Laminar ventricular myocardium on a microelectrode array-based chip. J Mater Chem B 2016;4:3534-43. [Crossref]
- Lind JU, Busbee TA, Valentine AD, et al. Instrumented cardiac microphysiological devices via multimaterial three-dimensional printing. Nat Mater 2016. [Epub ahead of print]. [Crossref] [PubMed]
- Wang G, McCain ML, Yang L, et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat Med 2014;20:616-23. [Crossref] [PubMed]


