
Tissue-specific tumour microenvironments are an emerging determinant of immunotherapy responses
Introduction
Cancer immunotherapies, which aim to enhance the anti-tumour immune response, have revolutionised the treatment of cancer. Various agents, known collectively as checkpoint blockade therapy, to block interactions of PD-1 and CTLA4 on T cells with their respective ligands, have been clinically approved in a variety of cancer types (1). Targeting these proteins removes the inhibitory effect on T cell function and restores effective anti-tumour responses (2). The results of checkpoint blockade have been particularly striking in melanoma and non-small cell lung cancer (NSCLC), which have achieved objective response rates of 40–45% (3,4). However, not all cancer types or patients benefit from these treatments and as such, patterns of response and resistance to checkpoint blockade therapies are under intense investigation. The current understanding is that additional mechanisms and cellular mediators also play roles in determining the outcome of checkpoint blockade therapies (5-7). In particular, the immunosuppressive tumour microenvironment (TME) is a major contributor to checkpoint blockade resistance (8).
During cancer progression, tumour cells manipulate the surrounding microenvironment to create an immunosuppressive, pro-tumourigenic niche. In this way tumours are like a complex organ comprised of various cell types, each performing certain functions. The key cellular mediators of the TME include regulatory T cells (Tregs), tumour associated macrophages (TAMs) and myeloid derived suppressor cells (MDSCs) (8). These cell types suppress anti-tumour immune responses via different mechanisms, including secreting immunosuppressive molecules such as IL-10, TGFβ, IDO and adenosine, and compete for resources with anti-tumour effector cells. Depleting these immunosuppressive cells or targeting immunosuppressive mediators leads to reduced tumour growth and enhanced response to checkpoint blockade (8).
Current evidence for tissue-specific TMEs
Further understanding of the immunosuppressive TME will undoubtedly improve immunotherapeutic outcomes. An understudied area in determining the composition of the TME is the anatomical location of tumour growth (9-11). Multiple murine studies have shown differences in the protein, RNA, metabolite and cellular composition of the TME depending on the location of the tumour (9). As a consequence, the same tumour cell line growing in different locations in mice results in varied responses to chemotherapy, anti-angiogenic therapy and immunotherapies including vaccination, checkpoint blockade and agonistic antibody regimes (9,12).
Studies of the TME in humans have also revealed tissue-specific TMEs and demonstrated correlations between certain metastatic locations and responses to checkpoint blockade in melanoma, NSCLC, triple negative breast cancer (TNBC) and urothelial cancer (13-15). For example, reduced responses to αPD-1 monotherapy and presence of liver metastases has been reported in patients with melanoma (13), urothelial cancer (15), TNBC (14) and NSCLC (13,16). In contrast, other studies have shown no association of checkpoint blockade response with the presence of liver metastases (9,13). Secondary lung metastases in NSCLC patients showed reduced response to αPD-1 in one patient cohort, but not in others (9,13). The influence of lung metastases in melanoma also differed between studies with some studies revealing a positive association and others revealing no association with checkpoint blockade responsiveness (9,13). However these results are complicated by the genetic heterogeneity of the human tumour cells (10) and differences in location, tumour type and the specific therapy used. Additionally, these associations are difficult to dissect since metastasis to specific organs, such as the liver, can be attributed to more advanced patients and larger tumour burden (16).
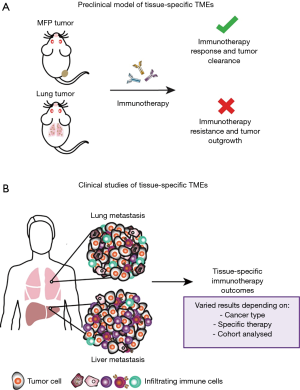
A recent study carried out in our laboratory aimed to enhance the understanding of the tissue-specific TME by utilising clinically relevant mouse models (Figure 1A) (17). We investigated breast cancer growth in common sites of metastasis, to extend comparisons of subcutaneous and orthotopic sites of tumour growth in previous studies (12), and how tumours growing in these sites influenced response to immunotherapies. Specifically, we focused on the 67NR murine breast cancer model, a non-metastatic cell line derived from the BALB/c syngeneic breast cancer cell line, 4T1, and compared tumours growing in the mammary fat pad (MFP), and the liver and lungs as common sites of metastasis. We found that tumours growing orthotopically in the MFP and in the liver were more responsive to αPD-1/αCTLA4 and an experimental immunotherapy known as trimAb (αDR5, α4-1BB and αCD40) than the same tumour line growing in the lungs. Similar findings were observed in the Renca kidney murine cancer cell line where tumours growing subcutaneously were more responsive than tumours growing in the lungs to trimAb. Consistent with our findings, Bialkowski et al. revealed that TC-1 lung tumours growing in the lungs were less responsive to a vaccine regime than subcutaneous tumours (18).
The difference in therapeutic responses was not due to inherent differences in the tumour cells. This was evidenced by assessment of therapy relevant proteins on tumour cells, such as PD-L1 and DR5, taken from the lung or MFP, and by reinjecting tumour cells derived from either site back to the same or opposite location. Additionally, we found no differences in vascularisation or compound permeability between lung and MFP tumours. Assessment of the expression profiles between lung, liver and MFP tumours revealed distinct immune gene expression profiles dependent on the tumour location. Thus, we hypothesised that the difference in immunotherapy responses was due to distinct differences in the immune TME of tumours growing in the lung or MFP.
Indeed, when we interrogated the TME using multiparameter flow cytometry, gene expression analysis, immunohistochemistry (IHC) and ex vivo stimulation of T cells and NK cells, we found that lung tumours had a more immunosuppressive TME than MFP tumours. Compared with MFP tumours, lung tumours had a lower percentage of CD8+ T cells and a higher percentage of Ly6C/Ly6G+ myeloid cells, which are markers for MDSCs, within the total immune infiltrate in these tumours. Furthermore, the responsive MFP tumours had significantly reduced infiltration of MDSC post αPD-1/αCTLA4 treatment, a phenomenon that was not observed in therapy resistant lung tumours. These observations are similar to the study that used the TC-1 tumour model, which revealed that resistant genital tract tumours had higher frequency of MDSCs that were not decreased by administration of a therapeutic vaccine (18). In another study using the CT26 model, comparison of the tumours growing at different locations, revealed that response to αPD-1/αCTLA4 was associated with increased presence of T cells and decreased MDSCs (19). Together, these studies highlight that the TME at different anatomical sites is a determining factor that influences the response to immunotherapy.
Interestingly, our study indicates that the same immune cell type can play divergent roles at different locations. Compared to NK cells isolated from MFP tumours, NK cells from lung tumours had decreased expression of CD69 and reduced ability to produce IFNγ when stimulated ex vivo. Although NK cells within the lungs were poorly activated at the timepoints analysed, NK cells still appeared to play an important role in the anti-tumour immune response in lung tumours compared to MFP tumours. We interrogated the contribution of various immune cell subsets following immunotherapy treatment using depleting antibodies against CD8, CD4, asialoGM1, F4/80 and Ly6G or GR1. Depletion of NK cells with anti-asialoGM1 resulted in limited reduction in therapeutic response of MFP tumours after immunotherapy but completed abrogated the anti-tumour effect in lung tumours. Across both tumour locations and therapies, CD8+ T cell depletion resulted in complete reversal of the therapeutic effect. Therefore, the immune cell subsets responsible for anti-tumour and therapy responses differed between MFP and lung tumours. In lung tumours, both NK and CD8+ T cells are the key drivers in response to these immunotherapies, while in the MFP, NK cells do not play a decisive role. Consistent with our findings, a study in the B16F10 model also provides evidence of tissue-specificity in the immune cell mediators important for inducing an effective anti-tumour response. Antibody-dependent cellular cytotoxicity (ADCC) by tumour directed antibodies was mediated by tissue-resident macrophages in B16F10 lung tumours however this occurred by a different subset of macrophages derived from Ly6C+ monocytes in B16F10 subcutaneous tumours (20).
Perspectives and future directions
To date, few studies have retrospectively analysed patients treated with checkpoint blockade for correlations between sites of metastasis and responsiveness (9). The majority of data has come from melanoma and NSCLC, which are among the first cancers FDA approved for checkpoint blockade treatment. However, in all studies, organ specific associations with checkpoint blockade response are varied and may be dependent on the cancer type, cohort analysed and specific therapy used (Figure 1B).
Our study in murine models identified specific differences in the TME depending on tumour location, and demonstrated that these differences are likely to be responsible for the disparate therapeutic efficacy observed. Given the dynamic nature of the TME this work is complex and unanswered questions from ours and other murine studies still remain. One of the most intriguing questions is how murine studies of tissue-specific TMEs correspond to immunotherapy responses and tumour locations in the clinical setting.
Future studies with a clear aim of understanding tissue-specific TMEs and immunotherapy responses in the human setting should be undertaken. Specifically, more detailed analysis of human tumours will instruct the development of more relevant murine models to better reflect the clinical setting for further mechanistic studies. Ideally, in depth analysis of the TME in primary and metastatic human tumours before, during and after the course of therapy along with detailed tracking of specific lesion growth over time is needed to enhance our knowledge of tissue-specific patterns of response. A case-study from Jiménez-Sànchez et al. provided detailed analysis of an ovarian cancer patient over the course of various chemotherapies (21). This study tracked tumour growth over a period of nine years in four metastatic locations including the liver, spleen, vaginal cuff and right upper quadrant (RUQ). The responding lesions, liver and RUQ metastases, were associated with a T cell inflamed TME with increased T cell signalling and T cell clonal expansion compared to T cells from the primary tumour and non-responding lesions, spleen and vaginal cuff metastases, which had increased Wnt signalling and immune exclusion. This level of tissue collection would allow insight into tumours that have failed therapy and those that have responded. However, given that biopsies and detailed imaging can be highly invasive for patients, expensive and time consuming, getting such detailed information is practically difficult.
A possible way around collecting invasive biopsies while still analysing metastatic sites is the establishment of rapid autopsy programs (22). In these programs, patients and their families consent to rapid tissue collection, within hours of death, and use of the patients’ tissues for research purposes. Such programs require collaborative effort of patients, families, clinicians, hospital staff, autopsy teams and researchers, and the tissues derived can have profound implications on research outcomes. This approach is highly beneficial in collecting large quantities of fresh tissue from multiple locations, especially locations that are challenging to biopsy in living patients. Tissues harvested from rapid autopsy have been used for DNA, RNA, proteomic and IHC analysis as well as establishing patient-derived xenografts in prostate, pancreatic, ovarian, breast and brain cancers (22). Although these tissues only include tumours that have failed therapy, the collection of tissue from multiple metastatic locations post-mortem provides valuable resource for understanding the tissue-specific differences in the immune microenvironment, especially if these samples are from immunotherapy treated patients.
Due to the valuable nature of tumour biopsies and post-mortem tissues, all efforts should be considered to maximise the information gained. Ideally, this means that tissues are utilised as fresh, frozen and/or paraffin embedded for various analyses including genomic, transcriptomic, proteomic, epigenetic, metabolomic, and immunologic studies. Fortunately, there is a plethora of high-throughput techniques that can capitalise on available tissue. In particular, single cell sequencing (23), mass cytometry (23) and multiplexing IHC (24) approaches have revolutionised immunologic analyses. Additionally, techniques such as spatial transcriptomics and in situ sequencing are able to combine these cutting-edge technologies to gain a high degree of information on a per cell basis including detailed phenotype and spatial information (25).
Focused analysis of existing samples (especially biobanked rapid autopsy samples) and directed collection of new samples for the purpose of understanding tissue-specific microenvironments in various metastatic sites will lead to valuable insights. In addition, advancement of clinical imaging techniques, such as utilising cell surface markers for in vivo imaging of T cell activation, could allow accurate detection of real time changes within the TME. Taking these findings back into murine models will allow manipulation of immunotherapy resistant tumour types to identify improvements to existing immunotherapies or development of novel immunotherapies. Furthermore, established tissue-specific response patterns can provide prognostic information and instruct therapeutic decision making. This second wave of immunotherapy research is vital in the continuation and improvement to initial clinical successes of the first wave of immunotherapies.
Acknowledgments
Funding: None.
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editors (Peng Luo, Clare Y. Slaney and Jian Zhang) for the series “Immunotherapy and Tumor Microenvironment” published in Journal of Thoracic Disease. The article did not undergo external peer review.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jtd.2020.03.64). The series “Immunotherapy and Tumor Microenvironment” was commissioned by the editorial office without any funding or sponsorship. CYS served as the unpaid Guest Editor of the series. The authors have no other conflicts of interest to declare.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Hargadon KM, Johnson CE, Williams CJ. Immune checkpoint blockade therapy for cancer: An overview of FDA-approved immune checkpoint inhibitors. Int Immunopharmacol 2018;62:29-39. [Crossref] [PubMed]
- Wei SC, Duffy CR, Allison JP. Fundamental Mechanisms of Immune Checkpoint Blockade Therapy. Cancer Discov 2018;8:1069-86. [Crossref] [PubMed]
- Garon EB, Rizvi NA, Hui R, et al. Pembrolizumab for the treatment of non-small-cell lung cancer. N Engl J Med 2015;372:2018-28. [Crossref] [PubMed]
- Larkin J, Chiarion-Sileni V, Gonzalez R, et al. Combined Nivolumab and Ipilimumab or Monotherapy in Untreated Melanoma. N Engl J Med 2015;373:23-34. [Crossref] [PubMed]
- Simpson TR, Li F, Montalvo-Ortiz W, et al. Fc-dependent depletion of tumor-infiltrating regulatory T cells co-defines the efficacy of anti-CTLA-4 therapy against melanoma. J Exp Med 2013;210:1695-710. [Crossref] [PubMed]
- Beavis PA, Henderson MA, Giuffrida L, et al. Dual PD-1 and CTLA-4 Checkpoint Blockade Promotes Antitumor Immune Responses through CD4(+)Foxp3(-) Cell-Mediated Modulation of CD103(+) Dendritic Cells. Cancer Immunol Res 2018;6:1069-81. [Crossref] [PubMed]
- House IG, Savas P, Lai J, et al. Macrophage derived CXCL9 and CXCL10 are required for anti-tumor immune responses following immune checkpoint blockade. Clinical Cancer Research 2019:clincanres.1868.2019.
- Binnewies M, Roberts EW, Kersten K, et al. Understanding the tumor immune microenvironment (TIME) for effective therapy. Nat Med 2018;24:541-50. [Crossref] [PubMed]
- Oliver AJ, Lau PKH, Unsworth AS, et al. Tissue-Dependent Tumor Microenvironments and Their Impact on Immunotherapy Responses. Front Immunol 2018;9:70. [Crossref] [PubMed]
- Salmon H, Remark R, Gnjatic S, et al. Host tissue determinants of tumour immunity. Nat Rev Cancer 2019;19:215-27. [PubMed]
- Horton BL, Fessenden TB, Spranger S. Tissue Site and the Cancer Immunity Cycle. Trends Cancer 2019;5:593-603. [Crossref] [PubMed]
- Devaud C, Westwood JA, John LB, et al. Tissues in different anatomical sites can sculpt and vary the tumor microenvironment to affect responses to therapy. Mol Ther 2014;22:18-27. [Crossref] [PubMed]
- Tumeh PC, Hellmann MD, Hamid O, et al. Liver Metastasis and Treatment Outcome with Anti-PD-1 Monoclonal Antibody in Patients with Melanoma and NSCLC. Cancer Immunol Res 2017;5:417-24. [Crossref] [PubMed]
- Loi S, Adams S, Schmid P, et al. LBA13Relationship between tumor infiltrating lymphocyte (TIL) levels and response to pembrolizumab (pembro) in metastatic triple-negative breast cancer (mTNBC): Results from KEYNOTE-086. Ann Oncol 2017;28:605-49. [Crossref]
- Zheng Y, Narwal R, Jin C, et al. Population Modeling of Tumor Kinetics and Overall Survival to Identify Prognostic and Predictive Biomarkers of Efficacy for Durvalumab in Patients With Urothelial Carcinoma. Clin Pharmacol Ther 2018;103:643-52. [Crossref] [PubMed]
- Shiroyama T, Suzuki H, Tamiya M, et al. Clinical Characteristics of Liver Metastasis in Nivolumab-treated Patients with Non-small Cell Lung Cancer. Anticancer Res 2018;38:4723-9. [Crossref] [PubMed]
- Oliver AJ, Davey AS, Keam SP, et al. Tissue-specific tumor microenvironments influence responses to immunotherapies. Clin Transl Immunology 2019;8:e1094. [Crossref] [PubMed]
- Bialkowski L, van Weijnen A, Van der Jeught K, et al. Intralymphatic mRNA vaccine induces CD8 T-cell responses that inhibit the growth of mucosally located tumours. Sci Rep 2016;6:22509. [Crossref] [PubMed]
- Zhao X, Li L, Starr TK, et al. Tumor location impacts immune response in mouse models of colon cancer. Oncotarget 2017;8:54775-87. [Crossref] [PubMed]
- Lehmann B, Biburger M, Bruckner C, et al. Tumor location determines tissue-specific recruitment of tumor-associated macrophages and antibody-dependent immunotherapy response. Sci Immunol 2017. [Crossref] [PubMed]
- Jiménez-Sánchez A, Memon D, Pourpe S, et al. Heterogeneous Tumor-Immune Microenvironments among Differentially Growing Metastases in an Ovarian Cancer Patient. Cell 2017;170:927-38.e20. [Crossref] [PubMed]
- Duregon E, Schneider J, DeMarzo AM, et al. Rapid research autopsy is a stealthy but growing contributor to cancer research. Cancer 2019;125:2915-9. [Crossref] [PubMed]
- Lyons YA, Wu SY, Overwijk WW, et al. Immune cell profiling in cancer: molecular approaches to cell-specific identification. NPJ Precis Oncol 2017;1:26. [Crossref] [PubMed]
- Halse H, Colebatch AJ, Petrone P, et al. Multiplex immunohistochemistry accurately defines the immune context of metastatic melanoma. Sci Rep 2018;8:11158. [Crossref] [PubMed]
- Rodriques SG, Stickels RR, Goeva A, et al. Slide-seq: A scalable technology for measuring genome-wide expression at high spatial resolution. Science 2019;363:1463-7. [Crossref] [PubMed]


