
Role of immunotherapy and co-mutations on KRAS-mutant non-small cell lung cancer survival
Introduction
Lung cancer remains the leading cause of cancer death in the world with 1.69 million deaths in 2015 (1). It is estimated that there will be 228,150 new cases and 142,670 deaths in 2019 (2). Lung cancers are divided into two major subtypes, small cell and non-small cell. Majority of lung cancers are non-small cell lung cancer (NSCLC) accounting for around 85% of all cases with small-cell making up most of the remaining at 15% (3). NSCLC is composed of many histologic subtypes with 40% of lung cancers are adenocarcinoma, in addition squamous cell (25%) and large cell (15%) being the most common. A few other subtypes including adenosquamous and sarcomatoid carcinoma make up the remaining. Direct and indirect exposure to tobacco smoke is the predominant risk factor. Other risk factors include residential radon, indoor air pollution, asbestos, and paint dust (4). There is also increasing interest in genetic susceptibility as one-fourth of all lung cancer patients are never smokers and the associated possibility of these patients harboring treatable oncogenic alterations (5).
The molecular era of oncology has changed the way lung cancer is treated, especially for NSCLC. NSCLC is a heterogeneous disease with variable molecular mutations. Vi-Ki-ras2 Kirsten rat sarcoma viral oncogene (KRAS) is one of the most common oncogenic drivers, especially in lung cancer, and is found in around 25–30% adenocarcinomas. The other molecular abnormalities related to RAS pathway are EGFR (10–23%), BRAF (2%), MET (2%), HER2 (1%) and NRAS (0.2%). As there are no targeted therapies for KRAS patients, the majority of the patients are typically treated with cytotoxic chemotherapy in combination with immunotherapy or immunotherapy alone. However, a retrospective study of 282 patients with advanced NSCLC treated with ICI compared the efficacy of ICI in patients with KRAS mutations versus without KRAS mutations (6). Jeanson et al. showed no significant differences in treatment outcomes for patients with KRAS mutations compared to those without KRAS mutations with similar overall response rate (ORR: 18.7% vs. 14.4%, P=0.348), progression-free survival (PFS: 3.09 vs. 2.66 months; P=0.584) and overall survival (OS: 14.29 vs. 11.14 months; P=0.682) (6). However, there was a trend towards improved ORR and prolonged PFS in patients with KRAS mutations and programmed death-ligand 1 (PD-L1) ≥50%, which was not observed in the non-KRAS mutant cohort (6). This and other studies (7-9) warrant further investigation into the role of immunotherapy for KRAS patients and the correlation with PD-L1 expression.
KRAS gene and cancer
KRAS is one of the most common gene mutations in hematologic and solid tumors. The behavior of KRAS is varied across malignancies and this requires different strategies to manage. The KRAS gene (chromosome 12p12.1) is primarily involved in regulating cell division. It is a member of the RAS family of genes that encodes four proteins that are highly related mediators of the mitogen-activated protein kinase (MAPK) pathway: HRAS, KRAS 4a, KRAS 4b and NRAS (10). These proteins function as guanosine triphosphatases (GTPases), binary switches that turn on and turn off multiple pathways involved in survival, proliferation, angiogenesis and differentiation via effector proteins. Activation of KRAS is controlled by binding to guanine triphosphate (GTP) and deactivation by guanine diphosphate (GDP) and its function is thus dependent on GTP/GDP ratio. GTPase activity is regulated through an interchange between GTPase activating proteins (GAPs) and guanine nucleotide exchange factors (GEFs) which control the ratio of active RAS-GTP and inactive RAS-GDP (11).
In the active GTP-bound state, the RAS family of proteins are involved in signaling of numerous downstream targets. The RAS/RAF/MEK/ERK is a pathway involved in regulation of the cell cycle and effecting other proliferation related proteins. The epidermal growth factor receptor (EGFR), fibroblast growth factor receptor (FGFR) and platelet-derived growth factor receptor (PDGF-R) among others are cell surface receptors that activate this pathway. RAS also promotes cell survival via PI3K/PDK1/AKT intracellular signaling. Tumor invasion and metastasis-inducing protein 1 (TIAM1), RALGDS and RALGDS-like proteins are involved in membrane trafficking. These downstream signaling pathways and others have been implicated in tumorigenesis.
Within KRAS, the most common mutations are G12C (40%), G12V (21%), G12D (17%), G12A (10%) and other (12%) G12 and G13 mutations (12). KRAS transversions (G-T, G-C) are typical for smokers and transitions (G-A) are typical for never smokers. KRAS mutations are associated with poorer outcomes in NSCLC (13). Renaud et al. showed that KRAS mutant patients had worse outcomes compared to wild type cases (13). Based off WHO classification system, there are five subtypes of adenocarcinoma (lepidic, acinar, papillary, micropapillary, solid). Lepidic invasive adenocarcinoma is divided into mucinous and non-mucinous types. Non-mucinous is associated with EGFR mutations whereas mucinous types are very commonly KRAS-mutated. These patients are often non-smokers. Mascaux et al. published a meta-analysis (14) showing a hazard ratio (HR) of 1.35 for patients that were KRAS-mutated with studies involving predominantly Asian populations confirmed these findings (15). In contrast, studies that included primarily western patients did not show that KRAS patients did worse as compared to KRAS wild type patients (15).
In these meta-analyses that had a poorer survival, it is postulated by Zer et al. that since KRAS mutants are generally mutually exclusive of EGFR mutants and Asian populations have higher percent of EGFR mutants (up to 40%) this selects for patients who are KRAS wild-type and possibly EGFR mutant, who have a better survival (16). KRAS mutations may have a weak association with worse prognosis (HR 1.3–1.5) though western population data does not support this and therefore has limited clinical utility (14). It also remains unclear if KRAS mutations are predictive of benefit for certain NSCLC patients receiving chemotherapy. A prospective study of 482 patients evaluated cisplatin and vinorelbine in the adjuvant setting in patients with NSCLC (17). Patients stratified by KRAS status suggests that KRAS mutant patients did not receive as much benefit from adjuvant chemotherapy compared to KRAS wild type patients. However, further interaction tests comparing HR were not statistically significant. In evaluation of KRAS mutations stratified by codon, a meta-analysis was performed on 1,543 patients in four adjuvant chemotherapy studies (18). There appeared to be a non-significant trend of benefit for KRAS wild type patients who received adjuvant chemotherapy as opposed to KRAS codon 12 mutated NSCLC patients. In contrast, those who harbored a codon 13 KRAS mutation performed poorly in comparison to KRAS wild-type and other KRAS mutants (P<0.001) (18).
A retrospective study involving 1,971 NSCLC patients with EGFR and KRAS mutations performed by Renaud and colleagues have shown that KRAS mutations may be predictive of resistance to radiation therapy (19). Identifying ways to target these KRAS mutations may lead to benefit for patients in combination with other traditional means of treatment. Concurrent mutations have recently been found to possibly play a prognostic role and may indicate if patients may be more responsive to therapy (20). The most common co-existing mutations are TP53 (39%), STK11 (30%), KEAP1 (24%), RBM10 (15%) and PTPRD (15%) (21). TP53 has been strongly associated with enhanced proliferation and STK11 has been associated with suppression of immune surveillance (22).
Methods
Objectives
We performed a retrospective single center clinical study to determine survival of patients with a diagnosis of NSCLC and a KRAS mutation. We sought to determine possible associations between KRAS status and other co-occurring mutations, as well as the relationship between KRAS status and immunotherapy.
Study conduct
We screened a prospectively collected, single institute, NSCLC molecular database for patients with KRAS mutation. Patients with KRAS mutations with metastatic disease who were treated between January 1st, 2009 and January 1st, 2016 were selected for this study.
Electronic medical records of the identified patients were reviewed by the study investigators to capture patient characteristics, tumor molecular profile, and patient outcome. Demographics included age, sex, race, date of birth, treatment history, and metastatic sites. Molecular profiling data included KRAS status and other concurrent mutation status. All molecular assays were performed by Clinical Laboratory Improvement Amendments (CLIA) certified assays. Imaging studies were reviewed to assess metastatic sites. The City of Hope (COH) institutional review board approved this retrospective study.
Statistical analyses
Fisher exact test and independent t-test were used to examine associations between categorical and continuous variables, respectively. Survival was estimated using the Kaplan-Meier method and differences in survival were evaluated via the log-rank test. Cox proportional hazards were employed to assess effects of specific factors on survival. Statistical analysis was performed with SPSS v.18.
Results
Patient characteristics and treatment
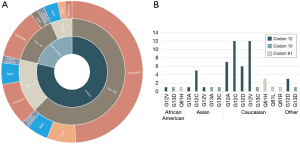
From 2009 to 2016, 60 patients with KRAS mutations were identified in the COH registry. Of these patients identified, 42 (70%) were stage IV, 7 (12%) stage I, 7 (12%) stage II, and 4 (7%) stage III at diagnosis. Forty-seven (78%) patients were smokers (former plus current). Caucasian was the most common (n=44, 73%) racial group, followed by Asian (n=9, 15%), African-American (n=3, 5%) and Pacific Islander (n=1, 1.7%). The average age at diagnosis was 67 (median 69.50) years; 30 patients (50%) were over 70 years, 23 (38%) patients were 51–69 years, and 7 (12%) 50 years or below. The most common histology was adenocarcinoma (n=52, 87%), followed by adenosquamous (n=3, 5%), large cell (n=2, 3%) and small cell, squamous cell and carcinosarcoma (n=1 each, less than 2% each). Majority of the patients had metastatic disease (n=52, 87%) with 20% (n=12) having brain metastasis. The average number of metastatic sites was 1.6 and patients received on average 1.97 (range, 0–5) lines of therapy including chemotherapy, biologic agents or immunotherapy. Twelve (20%) patients received immunotherapy with additional treatment modalities including radiation in 28 (47%) and surgery in 22 (37%) patients with a median OS at 15 months. Patient characteristics are shown in Figure 1A and Table 1.

Full table
Molecular status
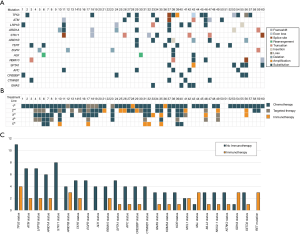
The most frequent molecular alteration was codon 12 mutation (n=47, 78%), followed by codon 13 (n=7, 12%) and codon 61 (n=6, 10%) mutations (Table 1). Patient KRAS mutations in each race is seen in Figure 1B. The most common co-occurring mutations in this cohort were TP53 (n=15, 25%), ATM (n=9, 15%), LRP1B (n=9, 15%), ARID1A (n=8, 13%), STK11 (n=8, 13%), ARID1B (n=7, 12%), TERT (n=7, 12%), EGFR (n=6, 10%), RBM10 (n=6, 10%), SPTA1 (n=6, 10%) (Figure 2A). Treatment characteristics are detailed in Table 1. In all 60 patients, systemic therapy given in up to 5 lines of treatment is shown in Figure 2B. The presence of co-occurring mutations in patients who were treated with immunotherapy versus patients who received no immunotherapy treatment is visualized in Figure 2C. RET mutated co-occurrence was the only unique instance in immunotherapy patients.
Survival
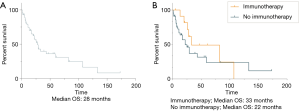
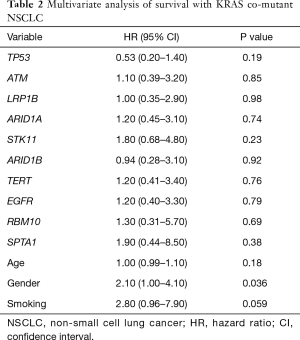
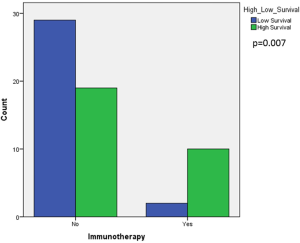
Median OS was 28 months in our patient cohort (Figure 3A). To evaluate the factors associated with likelihood of improved survival in this population, we focused on certain characteristics. There was an association with longer survival in patients who had an earlier stage, stages I, II, III, IV (58 vs. 26 vs. 3 vs. 11 months, respectively; P=0.002). In addition, there appeared to be a trend towards longer survival in those that received immunotherapy (33 months, n=12) in compared those who did not (22 months, n=48) (P=0.31) (Figure 3B). The most common co-occurring mutations in this cohort did not show a survival difference in comparison to patients who did not have those mutations: TP53 (P=0.019), ATM (P=0.85), LRP1B (P=0.99), ARID1A (P=0.74), STK11 (P=0.23), ARID1B (P=0.92), TERT (P=0.75), EGFR (P=0.79), RBM10 (P=0.69) and SPTA1 (P=0.38) (Table 2). Patients who harbored a codon 61 mutation had a median survival of 28 months compared to 11 months of codon 13 and 13 months of codon 12; this was not statistically significant (P=0.143). Based on the calculated median of survival (15 months; range, 1–173 months), we defined better survival as those patients who had a high survival (≥ median) and analyzed survival with those who received and did not receive immunotherapy. Our analysis revealed that there was an association between those patients who had high survival and who received immunotherapy (Figure S1, P=0.007).
Full table
Discussion
Lung cancer remains the number one cause of cancer death worldwide (1). KRAS mutations are among the most common molecular alterations identified in NSCLC. With decades of research, there are still no effective direct targets to the KRAS pathway. To better understand KRAS mutations, we evaluated co-mutations and identified factors associated with improved survival. The majority of KRAS mutations identified in our population were adenocarcinoma (87%), with small but similar distributions of other histologies including adenosquamous, large cell and small cell, squamous cell and carcinosarcoma, which is consistent with other previously published work of KRAS mutant NSCLC (23). In contrast to the Lung Adjuvant Cisplatin Evaluation (LACE)-Bio group who conducted a pooled analysis of patients enrolled in four randomized trials of adjuvant chemotherapy, we did not identify a trend towards patients being earlier in stage or younger (24). Instead, our patient population average age was 67 and majority were stage IV at diagnosis (70%).
Twenty-two percent of the patients in our cohort were never smokers which is much higher than previously reported distributions (25,26). In a series of KRAS-mutated lung adenocarcinoma where 17% of patients were never smokers, transition mutations were more common in never smokers (15%) compared to transversion mutations that were common in patients with smoking history (22%) (26). Our population had a similar distribution of transition mutations in smokers and never smokers, 26% and 31% respectively (P=0.705), and thus likely transition mutations are not the explanation as to why there is an increase in never smokers in our KRAS-mutated NSCLC population. Smoking-associated lung cancer differs among populations and 45–71% of Asian patients with lung cancer are never smokers (27,28). Our cohort of patients has an overrepresented Asian population (15%) in relation to 2010 US Census data (5.6%) and likely plays a role in the increase of KRAS never smokers in our cohort.
There remains continued interest in using KRAS as a prognostic marker in NSCLC. In support of previously published work (29,30), the most frequent KRAS mutation in our study was located on codon 12. Previous work by Yu et al. demonstrated that patients with KRAS codon 12 mutations had superior survival compared to those with codon 13 tumors, with a median of 16- and 13-month survival respectively (P=0.009) (29). We found a trend towards improved survival in patients with codon 61 mutations compared to codons 12 and 13, but this was not statically significant. This indicates that there are likely other additional biological factors that need further evaluation. Most recently STK11/LKB1 and KEAP1/NFE2L2 were found to be associated with primary resistance and worse outcomes (20,31).
We observed a number of unique co-mutations that have not been previously reported in KRAS mutants (20,30), such as RBM10 and SPTA1. RBM10 is a protein that binds RNA and functions by inhibiting proliferation of tumor cells and hence a tumor suppressor associated in regulation of Notch signaling (32). Defects in this gene are the cause of the X-linked recessive disorder, TARP syndrome that leads to several birth defects. RBM10 are frequently identified in adenocarcinomas of the lung and other cancer types including pancreatic, colorectal and thyroid. It is postulated that RBM10 is an RNA splicing regulator, once mutated leads to pathogenesis of adenocarcinoma due to deregulated splicing which can lead to proliferation (33). SPTA1 is a gene that encodes an actin crosslinking and molecular scaffold protein that links the plasma membrane to the actin cytoskeleton. Its exact nature and function in oncogenesis are unknown but it has been postulated to function as an oncogene (34). Mutations in this gene is associated with a hereditary red blood cell disorders including spherocytic hemolytic anemia, elliptocytosis type 2 and pyropoikilocytosis. It is a gene that is highly mutated in lung cancer (35). A retrospective study performed on 38 patients with small-cell lung cancer (SCLC) showed SPTA1 mutations expressed in all stages of SCLC and is thought to be associated with SCLC development (34).
Immunotherapy has recently become a vital therapeutic option in NSCLC as a first-line treatment alone or combined with chemotherapy (36,37). Twelve (20%) patients in our study received immunotherapy and we noted a correlation between patients who received immunotherapy and longer OS. However, a larger cohort analysis is necessary to evaluate the role of immunotherapy in KRAS-mutated patients. Ten of the 12 patients (83%) received immune checkpoint inhibitors as second-line or later lines of treatment. Despite this, there was a strong correlation between patients treated with immunotherapy and high survival. Previous studies have shown that KRAS mutations can induce PD-L1 overexpression through activation of the downstream pathways in NSCLC (7,38,39). Several co-occurring mutations, such as TP53 and LKB1, have been described as predictive biomarkers of clinical benefit—with TP53 co-mutations associated with clinical benefit while instances of LKB1 and KRAS mutants showed ineffectiveness of immunotherapy (40,41). The identification of these KRAS mutant subgroups may be the key towards identifying a biomarker of immunotherapy efficacy, as several recent studies demonstrated that KRAS mutation alone was not sufficient to predict immunotherapy response (6,42,43).
Effective treatments targeting KRAS mutations have represented a challenge so far. Checkpoint blockade has presented an intriguing area of study considering there has been limited advancement in additional cytotoxic therapies in the last few years. However, anti-PD-1 therapy has been an effective approach for KRAS mutated NSCLC patients without a validated biomarker (44). Most recent data show correlation of KRAS and high PD-L1 expression with improved outcomes (6,45). A large percentage of KRAS mutated NSCLC patients have positive smoking history (26). Tobacco-induced tumors present higher burden of mutation and neo-antigens. Higher neo-antigen burden was associated with improved PFS with anti-PD-1 therapy (44). Our findings are consistent with this in showing that there is a trend towards improved survival in KRAS mutant patients who received immunotherapy.
Ideally, prospective studies designed with interest in molecular alterations, prognosis, and ability for KRAS-mutated NSCLC to respond to various therapies would be helpful. Our study was limited to its retrospective design and limited selection of patients. Heterogeneity of treatment course in addition to next-generation sequencing (NGS) platforms used to analyze molecular alterations left our results not standardized. In conclusion, understanding the significance of co-mutations and their therapeutic implications, especially in response to immunotherapy and other agents represents an important step to develop better treatment options for KRAS-mutated lung cancers. Our findings warrant further investigation in a prospective setting with a larger data set.
Acknowledgments
We thank the clinical staff and nurses for their skill and dedication in treating and assisting the patients presented in this manuscript.
Funding: This work was supported by the National Cancer Institute of the National Institutes of Health under Grants P30CA033572, U54CA209978, R01CA218545.
Footnote
Provenance and Peer Review: This article was commissioned by the Guest Editor (Ammar Chaudhry) for the series “Role of Precision Imaging in Thoracic Disease” published in Journal of Thoracic Disease. The article was sent for external peer review organized by the Guest Editor and the editorial office.
Conflicts of Interest: All authors have completed the ICMJE uniform disclosure form (available at http://dx.doi.org/10.21037/jtd.2020.04.18). The authors have no conflicts of interest to declare. The series “Role of Precision Imaging in Thoracic Disease” was commissioned by the editorial office without any funding or sponsorship.
Ethical Statement: The authors are accountable for all aspects of the work in ensuring that questions related to the accuracy or integrity of any part of the work are appropriately investigated and resolved. The study was conducted in accordance with the Declaration of Helsinki (as revised in 2013). The study was approved by the City of Hope Institutional Review Board (No. 18433). Individual consent for this retrospective analysis was waived.
Open Access Statement: This is an Open Access article distributed in accordance with the Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License (CC BY-NC-ND 4.0), which permits the non-commercial replication and distribution of the article with the strict proviso that no changes or edits are made and the original work is properly cited (including links to both the formal publication through the relevant DOI and the license). See: https://creativecommons.org/licenses/by-nc-nd/4.0/.
References
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2017. CA Cancer J Clin 2017;67:7-30. [Crossref] [PubMed]
- Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin 2019;69:7-34. [Crossref] [PubMed]
- NCCN Clinical Practice Guidelines in Oncology. 2017.
- Torre LA, Siegel RL, Jemal A. Lung Cancer Statistics. Adv Exp Med Biol 2016;893:1-19. [Crossref] [PubMed]
- Parkin DM, Bray F, Ferlay J, et al. Global Cancer Statistics, 2002. CA Cancer J Clin 2005;55:74-108. [Crossref] [PubMed]
- Jeanson A, Tomasini P, Souquet-Bressand M, et al. Efficacy of Immune Checkpoint Inhibitors in KRAS-Mutant Non-Small Cell Lung Cancer (NSCLC). J Thorac Oncol 2019;14:1095-101. [Crossref] [PubMed]
- Chen N, Fang W, Lin Z, et al. KRAS mutation-induced upregulation of PD-L1 mediates immune escape in human lung adenocarcinoma. Cancer Immunol Immunother 2017;66:1175-87. [Crossref] [PubMed]
- Scheel AH, Ansen S, Schultheis AM, et al. PD-L1 expression in non-small cell lung cancer: Correlations with genetic alterations. Oncoimmunology 2016;5:e1131379. [Crossref] [PubMed]
- Dietrich M, Hunis B, Raez L. P2.07-052 Detection of KRAS Mutation in Blood Predicts Favorable Response to Immunotherapy in NSCLC. J Thorac Oncol 2017;12:S2149. [Crossref]
- Barbacid M. ras genes. Annu Rev Biochem 1987;56:779-827. [Crossref] [PubMed]
- Satoh T, Nakafuku M, Kaziro Y. Function of Ras as a molecular switch in signal transduction. J Biol Chem 1992;267:24149-52. [PubMed]
- Dogan S, Shen R, Ang DC, et al. Molecular epidemiology of EGFR and KRAS mutations in 3,026 lung adenocarcinomas: higher susceptibility of women to smoking-related KRAS-mutant cancers. Clin Cancer Res 2012;18:6169-77. [Crossref] [PubMed]
- Renaud S, Falcoz PE, Schaeffer M, et al. Prognostic value of the KRAS G12V mutation in 841 surgically resected Caucasian lung adenocarcinoma cases. Br J Cancer 2015;113:1206-15. [Crossref] [PubMed]
- Mascaux C, Iannino N, Martin B, et al. The role of RAS oncogene in survival of patients with lung cancer: a systematic review of the literature with meta-analysis. Br J Cancer 2005;92:131-9. [Crossref] [PubMed]
- Karachaliou N, Mayo C, Costa C, et al. KRAS mutations in lung cancer. Clin Lung Cancer 2013;14:205-14. [Crossref] [PubMed]
- Zer A, Ding K, Lee SM, et al. Pooled Analysis of the Prognostic and Predictive Value of KRAS Mutation Status and Mutation Subtype in Patients with Non–Small Cell Lung Cancer Treated with Epidermal Growth Factor Receptor Tyrosine Kinase Inhibitors. J Thorac Oncol 2016;11:312-23. [Crossref] [PubMed]
- Winton T, Livingston R, Johnson D, et al. Vinorelbine plus Cisplatin vs. Observation in Resected Non–Small-Cell Lung Cancer. N Engl J Med 2005;352:2589-97. [Crossref] [PubMed]
- Shepherd FA, Domerg C, Hainaut P, et al. Pooled Analysis of the Prognostic and Predictive Effects of KRAS Mutation Status and KRAS Mutation Subtype in Early-Stage Resected Non–Small-Cell Lung Cancer in Four Trials of Adjuvant Chemotherapy. J Clin Oncol 2013;31:2173-81. [Crossref] [PubMed]
- Renaud S, Schaeffer M, Voegeli AC, et al. Impact of EGFR mutations and KRAS amino acid substitution on the response to radiotherapy for brain metastasis of non-small-cell lung cancer. Future Oncol 2016;12:59-70. [Crossref] [PubMed]
- Arbour KC, Jordan E, Kim HR, et al. Effects of Co-occurring Genomic Alterations on Outcomes in Patients with KRAS-Mutant Non-Small Cell Lung Cancer. Clin Cancer Res 2018;24:334-40. [Crossref] [PubMed]
- Riely GJ, Jordan E, Kim HR, et al. Association of outcomes and co-occuring genomic alterations in patients with KRAS-mutant non-small cell lung cancer. J Clin Oncol 2016;34:9019. [Crossref]
- Schabath MB, Welsh EA, Fulp WJ, et al. Differential association of STK11 and TP53 with KRAS mutation-associated gene expression, proliferation and immune surveillance in lung adenocarcinoma. Oncogene 2016;35:3209-16. [Crossref] [PubMed]
- Guan JL, Zhong WZ, An SJ, et al. KRAS mutation in patients with lung cancer: a predictor for poor prognosis but not for EGFR-TKIs or chemotherapy. Ann Surg Oncol 2013;20:1381-8. [Crossref] [PubMed]
- Tsao M, Hainaut P, Bourredjem A, et al. 156O LACE-Bio pooled analysis of the prognostic and predictive value of KRAS mutation in completely resected non-small cell lung cancer (NSCLC). Ann Oncol 2010;21:viii63-77.
- Eberhard DA, Johnson BE, Amler LC, et al. Mutations in the epidermal growth factor receptor and in KRAS are predictive and prognostic indicators in patients with non–small-cell lung cancer treated with chemotherapy alone and in combination with erlotinib. J Clin Oncol 2005;23:5900-9. [Crossref] [PubMed]
- Riely GJ, Kris MG, Rosenbaum D, et al. Frequency and distinctive spectrum of KRAS mutations in never smokers with lung adenocarcinoma. Clin Cancer Res 2008;14:5731-4. [Crossref] [PubMed]
- Ou SH, Ziogas A, Zell JA. Asian ethnicity is a favorable prognostic factor for overall survival in non-small cell lung cancer (NSCLC) and is independent of smoking status. J Thorac Oncol 2009;4:1083-93. [Crossref] [PubMed]
- Gomez SL, Chang ET, Shema SJ, et al. Survival following non-small cell lung cancer among Asian/Pacific Islander, Latina, and Non-Hispanic white women who have never smoked. Cancer Epidemiol Biomarkers Prev 2011;20:545-54. [Crossref] [PubMed]
- Yu HA, Sima CS, Shen R, et al. Comparison of the characteristics and clinical course of 677 patients with metastatic lung cancers with mutations in KRAS codons 12 and 13. J Clin Oncol 2013;31:8025. [Crossref]
- Wood K, Hensing T, Malik R, et al. Prognostic and Predictive Value in KRAS in Non-Small-Cell Lung Cancer: A Review. JAMA Oncol 2016;2:805-12. [Crossref] [PubMed]
- Skoulidis F, Goldberg ME, Greenawalt DM, et al. STK11/LKB1 Mutations and PD-1 Inhibitor Resistance in KRAS-Mutant Lung Adenocarcinoma. Cancer Discov 2018;8:822-35. [Crossref] [PubMed]
- Hernandez J, Bechara E, Schlesinger D, et al. Tumor suppressor properties of the splicing regulatory factor RBM10. RNA Biol 2016;13:466-72. [Crossref] [PubMed]
- Zhao J, Sun Y, Huang Y, et al. Functional analysis reveals that RBM10 mutations contribute to lung adenocarcinoma pathogenesis by deregulating splicing. Sci Rep 2017;7:40488. [Crossref] [PubMed]
- Iwakawa R, Kohno T, Totoki Y, et al. Expression and clinical significance of genes frequently mutated in small cell lung cancers defined by whole exome/RNA sequencing. Carcinogenesis 2015;36:616-21. [Crossref] [PubMed]
- Carr TH, McEwen R, Dougherty B, et al. Defining actionable mutations for oncology therapeutic development. Nat Rev Cancer 2016;16:319-29. [Crossref] [PubMed]
- Mok TSK, Wu YL, Kudaba I, et al. Pembrolizumab versus chemotherapy for previously untreated, PD-L1-expressing, locally advanced or metastatic non-small-cell lung cancer (KEYNOTE-042): a randomised, open-label, controlled, phase 3 trial. Lancet 2019;393:1819-30. [Crossref] [PubMed]
- Gandhi L, Rodriguez-Abreu D, Gadgeel S, et al. Pembrolizumab plus Chemotherapy in Metastatic Non-Small-Cell Lung Cancer. N Engl J Med 2018;378:2078-92. [Crossref] [PubMed]
- Sumimoto H, Takano A, Teramoto K, et al. RAS-Mitogen-Activated Protein Kinase Signal Is Required for Enhanced PD-L1 Expression in Human Lung Cancers. PLoS One 2016;11:e0166626. [Crossref] [PubMed]
- Miura Y, Sunaga N, Kaira K, et al. Abstract 4028: Oncogenic KRAS mutations induce PD-L1 overexpression through MAPK pathway activation in non-small cell lung cancer cells. In: Proceedings of the 107th Annual Meeting of the American Association for Cancer Research; 2016 Apr 16-20; New Orleans, LA. Philadelphia (PA): AACR; Cancer Res 2016;76:Abstract nr 4028.
- Dong ZY, Zhong WZ, Zhang XC, et al. Potential Predictive Value of TP53 and KRAS Mutation Status for Response to PD-1 Blockade Immunotherapy in Lung Adenocarcinoma. Clin Cancer Res 2017;23:3012-24. [Crossref] [PubMed]
- Koyama S, Akbay EA, Li YY, et al. STK11/LKB1 Deficiency Promotes Neutrophil Recruitment and Proinflammatory Cytokine Production to Suppress T-cell Activity in the Lung Tumor Microenvironment. Cancer Res 2016;76:999-1008. [Crossref] [PubMed]
- Passiglia F, Cappuzzo F, Alabiso O, et al. Efficacy of nivolumab in pre-treated non-small-cell lung cancer patients harbouring KRAS mutations. Br J Cancer 2019;120:57-62. [Crossref] [PubMed]
- Lee CK, Man J, Lord S, et al. Clinical and Molecular Characteristics Associated With Survival Among Patients Treated With Checkpoint Inhibitors for Advanced Non-Small Cell Lung Carcinoma: A Systematic Review and Meta-analysis. JAMA Oncol 2018;4:210-6. [Crossref] [PubMed]
- Rizvi NA, Hellmann MD, Snyder A, et al. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 2015;348:124-8. [Crossref] [PubMed]
- Kim JH, Kim HS, Kim BJ. Prognostic value of KRAS mutation in advanced non-small-cell lung cancer treated with immune checkpoint inhibitors: A meta-analysis and review. Oncotarget 2017;8:48248-52. [Crossref] [PubMed]


